Introduction

About mefikit
mefikit — short for Mesh and Field Kit — is a comprehensive library for
generating, manipulating, and analyzing unstructured meshes together with
associated scalar, vector, and tensor fields. Its goal is to provide a unified
in-memory representation of meshes and fields, along with a set of robust,
efficient tools that support numerical simulation workflows in research,
engineering, and scientific computing.
mefikit focuses on three core principles:
- A flexible mesh model capable of representing mixed-element unstructured meshes.
- A consistent field and group architecture that attaches data to mesh entities across dimensions.
- Zero-copy, iterator-based access patterns to efficiently navigate and process large meshes.
This design positions mefikit as a modern core library for algorithm
development.
mefikit provides
umeshthe unstructured mesh container supporting mixed element types, fields, and groups.iomodules for reading and writing meshes in various formats (e.g., VTK, serde_json, serde_yaml).topologytools for analyzing mesh connectivity, computing descending meshes, neighbours, domain frontier, etc.geometrytools for computing element measures, centroids, etc.Selectorutilities for querying and filtering mesh elements based on geometric or topological criteria.
Definitions
Element
An element is an abstract mesh entity specified by:
- its type (e.g., triangle, hexahedron, polygon, polyhedron)
- its connectivity (indices referencing the global coordinate array)
In mefikit, elements are ephemeral, zero-copy views constructed on the fly
when iterating through a mesh block. They do not own memory; instead they refer
to underlying index and coordinate buffers. This behavior mirrors VTK’s “cell
iterator” pattern while improving cache locality and reducing overhead.
It is only accessible through the rust API. The python bindings do not expose
elements directly because python is only intended to be used for high-level
scripting. Do not fear of going to the rust side for performance critical code,
mefikit was designed to be simple to use for rust newcommers.
Topological Dimension
The topological dimension of an element is the dimension of the mathematical object it represents:
- 0D → vertices
- 1D → edges / segments
- 2D → faces
- 3D → volumes
This categorization is independent of the embedding space and is central for field association and mesh validity checks.
Spatial Dimension
The spatial dimension is the dimension of the coordinate system in which the mesh is embedded, typically 2D or 3D. All element coordinates must be consistent with this spatial dimension. For example, one may have:
- 2D elements in 2D (pure surface mesh)
- 2D elements in 3D (surface embedded in 3D)
- 3D volumetric elements in 3D
mefikit does not assume a fixed coordinate system beyond this requirement.
Mesh
A mesh in mefikit is a container that holds:
- the global node coordinate array,
- a set of blocks, each containing elements of a given type,
- fields (scalar/vector/tensor) attached to elements of a given topological dimension,
- groups, i.e., labeled subsets of elements for boundary conditions, materials, or post-processing.
Properties:
- You may attach any number of fields or groups.
- Groups may overlap arbitrarily.
- A field must cover all elements of its associated topological dimension (similar to Gmsh’s “node data” or “element data” consistency).
If these constraints seem restrictive, mefikit encourages using multiple
meshes (e.g., one per partition, or one per physical domain) to better match
specialized workflows.
Topologically Valid Mesh
A topologically valid mesh in mefikit satisfies the following conditions:
-
No overlapping higher-dimensional elements Two 3D elements may share faces or edges, but they must not occupy the same region of space. (Duplicate faces in a hex-dominant mesh are allowed and interpreted in context.)
-
Lower-dimensional entities derive consistently from higher-dimensional ones
- All edges must be edges of some face or volume.
- All faces must be boundaries of some volume.
- No “floating” or orphaned lower-dimension elements are allowed. This is consistent with the expectations of many finite-element and finite-volume codes.
-
Non-degenerate geometry Elements must not be geometrically degenerate:
- triangles must have non-zero area,
- tetrahedra must have non-zero volume,
- polygons must be well-defined and non-self-intersecting, etc. These checks are similar to the geometric validation steps performed by MeshGems and various meshing libraries.
mefikit Mesh Format
The mefikit format defines a compact, extensible, and expressive way to
represent unstructured meshes composed of heterogeneous elements — including
vertices, edges, faces, and volumetric cells. Supported element types include,
but are not limited to:
- 0D: vertices
- 1D: segments, polylines
- 2D: triangles, quadrilaterals, polygons
- 3D: tetrahedra, hexahedra, prisms, pyramids, polyhedra
In addition, any element can carry fields (scalar, vector, or tensor) that reside directly on its topological dimension. This enables workflows akin to VTK field data. Fields must be defined upon all elements of a given topological dimension.
Topological basic operations
Subgraph and neighbours computation
The subgraph computation is done supposing that the mesh is topologically valid. The subgraph of a volume mesh is defined by a set of either faces, edges or vertices elements depending on the subgraph relative dimension.
The subgraph computation and the neighbours computation are tightly linked. The neighbours are defined as the elements that share a common lower dimensional element. For example, two volumes are neighbours if they share a common face, two faces are neighbours if they share a common edge, and so on.
In mefikit, the neighbours computation builds an adjacency graph, having as
nodes the elements of the mesh and as edges the shared lower dimensional
elements. Then, the neighbours of an element can be found by looking at its
adjacent nodes in the graph.
Connectivity equivalence
There is different ways to represent the connectivity of an element. Here are the different class of equivalence:
- exact representation equality: two elements are equivalent if their connectivity is exactly the same, including the order of the nodes.
- rotational equivalence: two elements are equivalent if their connectivity can be made the same by rotating the order of the nodes. The topological shape is strictly equivalent under all operations, preserving the measure orientation.
- chiral equivalence: two elements are equivalent if their connectivity can be made the same by reversing the order of the nodes and rotating the nodes. This equivalence does not preserve the measure orientation (if it is positive or negative).
- node set equivalence: two elements are equivalent if they have the same set of nodes, regardless of the order. This equivalence is theoretical only, as it does not preserve the shape of the element. But it is very useful to detect duplicate elements in a mesh.
Python Examples
UMesh basics
import mefikit as mf
import numpy as np
import pyvista as pv
pv.set_plot_theme("dark")
pv.set_jupyter_backend("static")
Building cartesian meshes
volumes = mf.build_cmesh(
range(2), np.linspace(0.0, 1.0, 3), np.logspace(0.0, 1.0, 4) / 10.0
)
volumes
<UMesh at 0x7474980e7750>
print(volumes)
UMeshBase {
coords: [[0.0, 0.0, 0.1],
[1.0, 0.0, 0.1],
[0.0, 0.5, 0.1],
[1.0, 0.5, 0.1],
[0.0, 1.0, 0.1],
[1.0, 1.0, 0.1],
[0.0, 0.0, 0.2154434690031884],
[1.0, 0.0, 0.2154434690031884],
[0.0, 0.5, 0.2154434690031884],
[1.0, 0.5, 0.2154434690031884],
[0.0, 1.0, 0.2154434690031884],
[1.0, 1.0, 0.2154434690031884],
[0.0, 0.0, 0.46415888336127786],
[1.0, 0.0, 0.46415888336127786],
[0.0, 0.5, 0.46415888336127786],
[1.0, 0.5, 0.46415888336127786],
[0.0, 1.0, 0.46415888336127786],
[1.0, 1.0, 0.46415888336127786],
[0.0, 0.0, 1.0],
[1.0, 0.0, 1.0],
[0.0, 0.5, 1.0],
[1.0, 0.5, 1.0],
[0.0, 1.0, 1.0],
[1.0, 1.0, 1.0]], shape=[24, 3], strides=[3, 1], layout=Cc (0x5), const ndim=2,
element_blocks: {
HEX8: ElementBlockBase {
cell_type: HEX8,
connectivity: Regular(
[[0, 1, 3, 2, 6, 7, 9, 8],
[2, 3, 5, 4, 8, 9, 11, 10],
[6, 7, 9, 8, 12, 13, 15, 14],
[8, 9, 11, 10, 14, 15, 17, 16],
[12, 13, 15, 14, 18, 19, 21, 20],
[14, 15, 17, 16, 20, 21, 23, 22]], shape=[6, 8], strides=[8, 1], layout=Cc (0x5), const ndim=2,
),
fields: {},
families: [0, 0, 0, 0, 0, 0], shape=[6], strides=[1], layout=CFcf (0xf), const ndim=1,
groups: {},
},
},
}
The mesh is composed of a coordinates array, and several blocks.
volumes.to_pyvista().plot(show_edges=True)
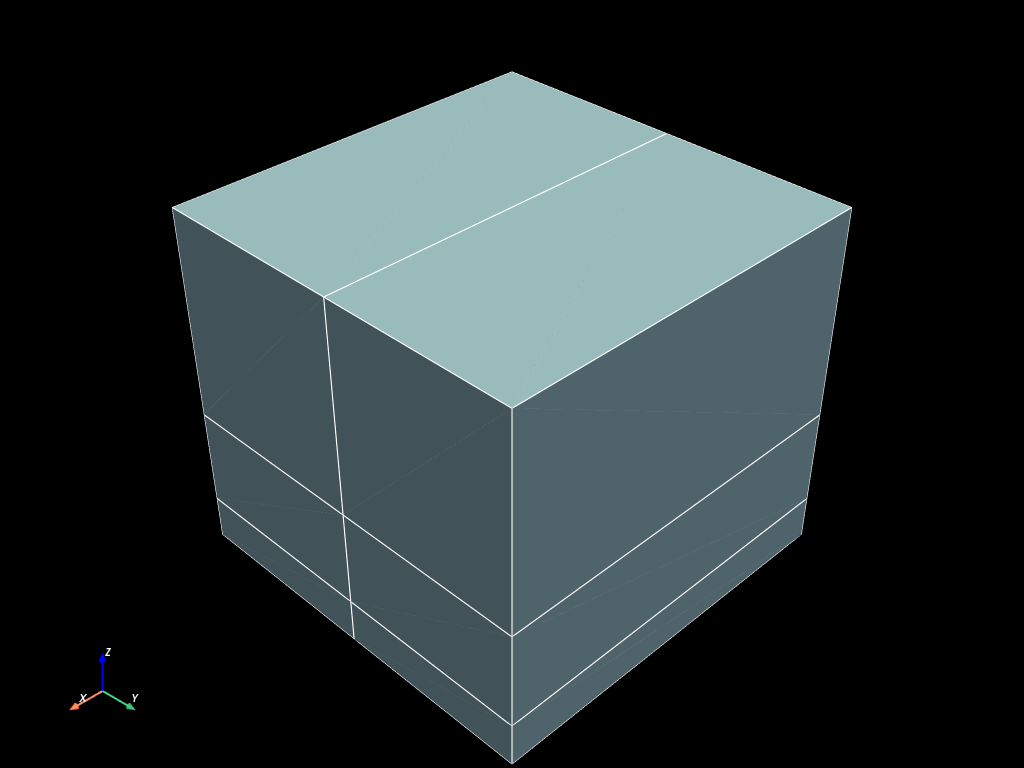

Building mesh with custom connectivity
x, y = np.meshgrid(np.linspace(0.0, 1.0, 5), np.linspace(0.0, 1.0, 5))
coords = np.c_[x.flatten(), y.flatten()]
conn = np.array(
[
[0, 1],
[1, 6],
[6, 5],
[5, 0],
[6, 7],
[7, 12],
[12, 11],
[11, 6],
[12, 17],
[17, 16],
[16, 11],
],
dtype=np.uint,
)
mesh = mf.UMesh(coords)
mesh.add_regular_block("VERTEX", np.arange(13, 22, dtype=np.uint)[..., np.newaxis])
mesh.add_regular_block("SEG2", conn)
mesh.add_regular_block("QUAD4", np.array([[3, 4, 9, 8]], dtype=np.uint))
mesh.to_pyvista(dim="all").plot(cpos="xy", show_edges=True)
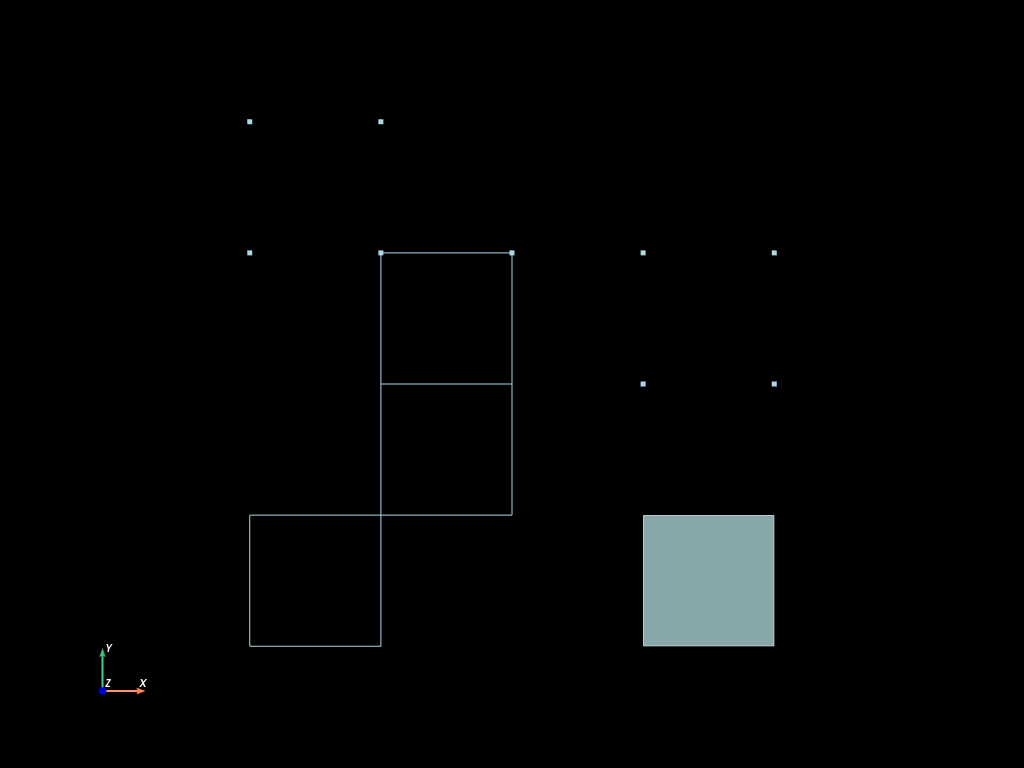
Mesh and Fields
x = np.logspace(-5, 0.0)
mesh2 = mf.build_cmesh(x, x)
mes = mesh2.measure()
mes
{'QUAD4': array([7.01482859e-12, 8.87274254e-12, 1.12227347e-11, ...,
2.74065140e-02, 3.46652722e-02, 4.38465503e-02], shape=(2401,))}
mesh2.measure_update()
mesh2.to_pyvista().plot()
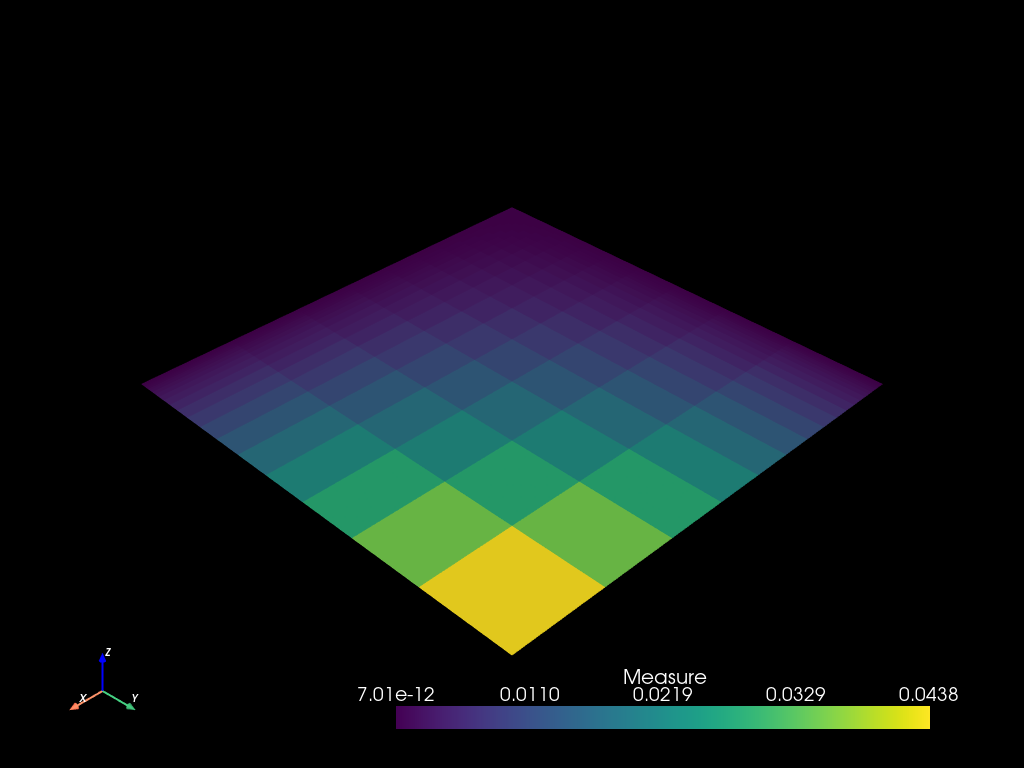
Intput / Output
import mefikit as mf
import numpy as np
volumes = mf.build_cmesh(
range(2), np.linspace(0.0, 1.0, 5), np.logspace(0.0, 1.0, 5) / 10.0
)
Memory exports
- Through numpy arrays manipulations:
- medcoupling
- meshio
- pyvista
- Through
stringtranslation toPython:- json
volumes.to_mc()
MEDCouplingUMesh C++ instance at 0x2e9f50f0. Name : "mf_UMesh". Not set !
volumes.to_pyvista()
| UnstructuredGrid | Information |
|---|---|
| N Cells | 16 |
| N Points | 50 |
| X Bounds | 0.000e+00, 1.000e+00 |
| Y Bounds | 0.000e+00, 1.000e+00 |
| Z Bounds | 1.000e-01, 1.000e+00 |
| N Arrays | 0 |
volumes.to_meshio()
<meshio mesh object>
Number of points: 50
Number of cells:
hexahedron: 16
volumes.to_json()
'{"coords":{"v":1,"dim":[50,3],"data":[0.0,0.0,0.1,1.0,0.0,0.1,0.0,0.25,0.1,1.0,0.25,0.1,0.0,0.5,0.1,1.0,0.5,0.1,0.0,0.75,0.1,1.0,0.75,0.1,0.0,1.0,0.1,1.0,1.0,0.1,0.0,0.0,0.17782794100389226,1.0,0.0,0.17782794100389226,0.0,0.25,0.17782794100389226,1.0,0.25,0.17782794100389226,0.0,0.5,0.17782794100389226,1.0,0.5,0.17782794100389226,0.0,0.75,0.17782794100389226,1.0,0.75,0.17782794100389226,0.0,1.0,0.17782794100389226,1.0,1.0,0.17782794100389226,0.0,0.0,0.31622776601683794,1.0,0.0,0.31622776601683794,0.0,0.25,0.31622776601683794,1.0,0.25,0.31622776601683794,0.0,0.5,0.31622776601683794,1.0,0.5,0.31622776601683794,0.0,0.75,0.31622776601683794,1.0,0.75,0.31622776601683794,0.0,1.0,0.31622776601683794,1.0,1.0,0.31622776601683794,0.0,0.0,0.5623413251903491,1.0,0.0,0.5623413251903491,0.0,0.25,0.5623413251903491,1.0,0.25,0.5623413251903491,0.0,0.5,0.5623413251903491,1.0,0.5,0.5623413251903491,0.0,0.75,0.5623413251903491,1.0,0.75,0.5623413251903491,0.0,1.0,0.5623413251903491,1.0,1.0,0.5623413251903491,0.0,0.0,1.0,1.0,0.0,1.0,0.0,0.25,1.0,1.0,0.25,1.0,0.0,0.5,1.0,1.0,0.5,1.0,0.0,0.75,1.0,1.0,0.75,1.0,0.0,1.0,1.0,1.0,1.0,1.0]},"element_blocks":{"HEX8":{"cell_type":"HEX8","connectivity":{"Regular":{"v":1,"dim":[16,8],"data":[0,1,3,2,10,11,13,12,2,3,5,4,12,13,15,14,4,5,7,6,14,15,17,16,6,7,9,8,16,17,19,18,10,11,13,12,20,21,23,22,12,13,15,14,22,23,25,24,14,15,17,16,24,25,27,26,16,17,19,18,26,27,29,28,20,21,23,22,30,31,33,32,22,23,25,24,32,33,35,34,24,25,27,26,34,35,37,36,26,27,29,28,36,37,39,38,30,31,33,32,40,41,43,42,32,33,35,34,42,43,45,44,34,35,37,36,44,45,47,46,36,37,39,38,46,47,49,48]}},"fields":{},"families":{"v":1,"dim":[16],"data":[0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0]},"groups":{}}}}'
File read/write
- On rust side, file I/O with the
read/writemethods:- vtk
- yaml
- json
Note that for now the vtk reader/writer is only based on the binary vtk 2.0 file format (which is quite old). No rust crate is doing better. I am planning on implementing a HDF5/CGNS compliant rust reader/writer (based on the hdf5 lib) in order to support a more HPC file format.
for ext in ("vtk", "yaml", "json"):
volumes.write(f"data/volumes.{ext}")
volumes_from_disk = mf.UMesh.read("data/volumes.vtk")
assert volumes_from_disk
assert (
volumes != volumes_from_disk
) # this is a new instance, with a different memory adress
Mesh extrusions
import mefikit as mf
import numpy as np
import pyvista as pv
pv.set_plot_theme("dark")
pv.set_jupyter_backend("static")
Building mesh with custom connectivity
First let’s build a 2D mesh wich will by used to demonstrate extrusions.
x, y = np.meshgrid(np.linspace(0.0, 1.0, 5), np.linspace(0.0, 1.0, 5))
coords = np.c_[x.flatten(), y.flatten()]
conn = np.array(
[
[0, 1],
[1, 6],
[6, 5],
[5, 0],
[6, 7],
[7, 12],
[12, 11],
[11, 6],
[12, 17],
[17, 16],
[16, 11],
],
dtype=np.uint,
)
mesh = mf.UMesh(coords)
mesh.add_regular_block("VERTEX", np.arange(13, 22, dtype=np.uint)[..., np.newaxis])
mesh.add_regular_block("SEG2", conn)
mesh.add_regular_block("QUAD4", np.array([[3, 4, 9, 8]], dtype=np.uint))
mesh.to_pyvista(dim="all").plot(cpos="xy", show_edges=True)
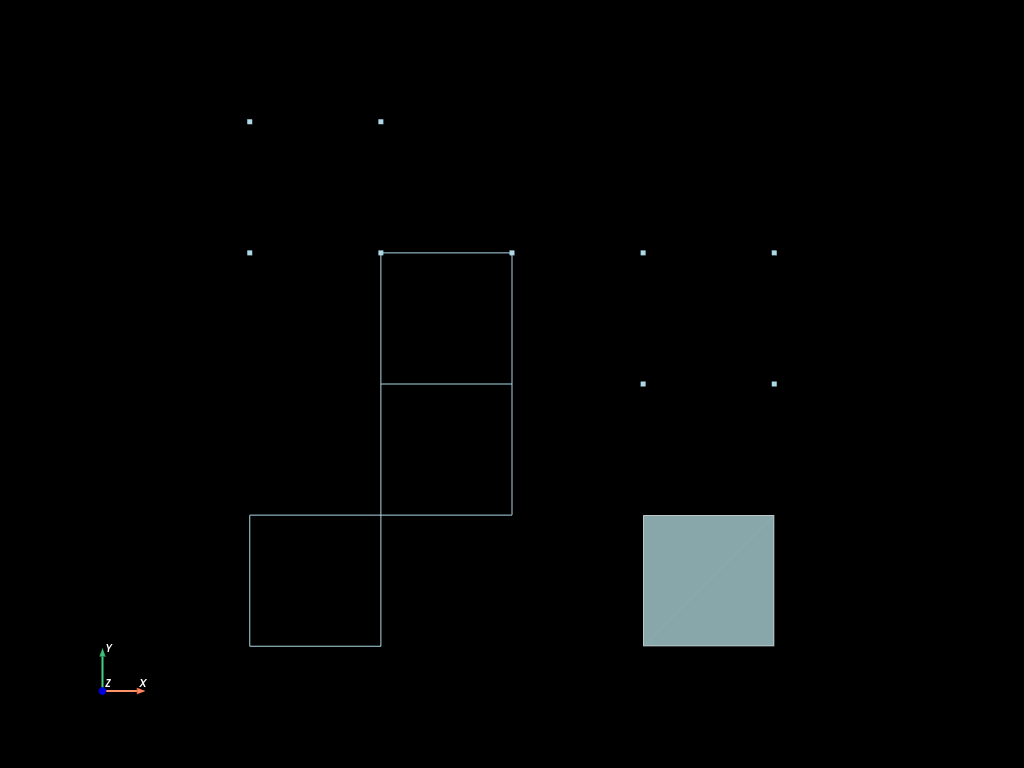
Extrusion along an existing axis
Build simple extruded mesh
extruded = mesh.extrude(range(3))
extruded.to_pyvista(dim="all").plot(show_edges=True)
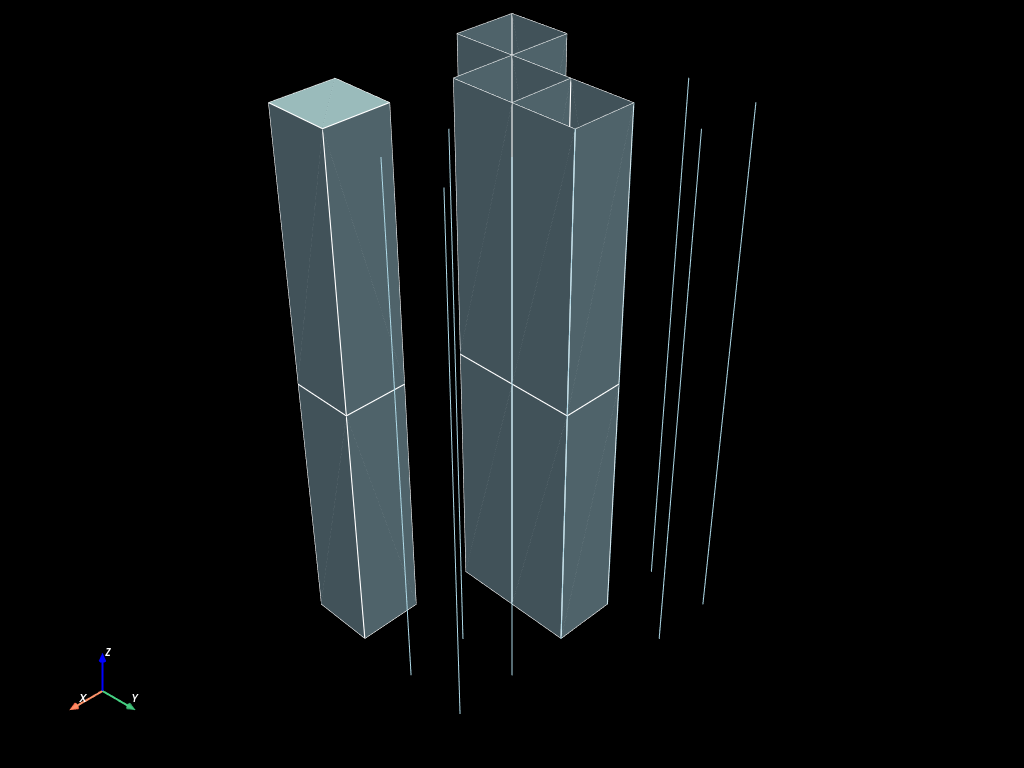

Build extruded mesh along a 3d line with parallel z faces
n = 50
x = np.sin(np.linspace(0.0, np.pi, n))
y = np.cos(np.linspace(0.0, np.pi, n))
z = np.linspace(0.0, 4.0, n)
line = np.c_[x, y, z]
extruded_par = mesh.extrude_parallel(line)
extruded_par.to_pyvista(dim="all").plot(show_edges=True)
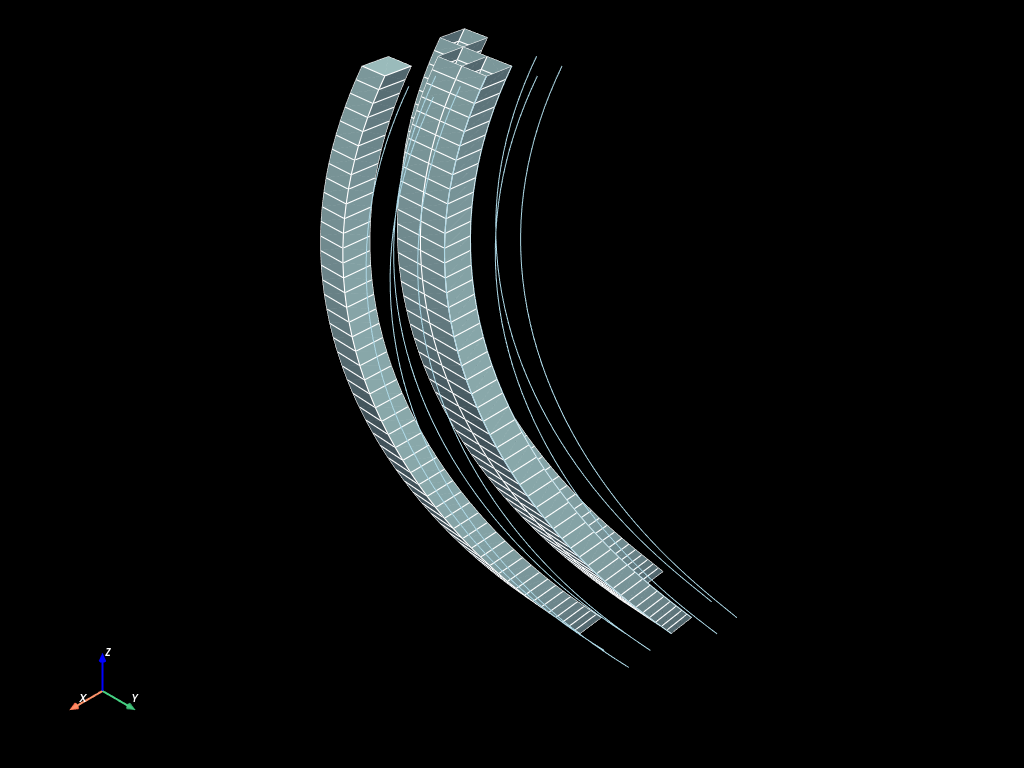

Build curviliear extrusion mesh
mesh = mf.build_cmesh(range(2), range(2))
n = 20
# x = np.linspace(0, 4, n)
x = np.zeros((n,))
# x = np.sin(np.linspace(0.0, np.pi, n))
y = np.cos(np.linspace(0.0, np.pi, n))
z = np.sin(np.linspace(0.0, np.pi, n))
line = np.c_[x, y, z]
extruded_curv = mesh.extrude_curv(line)
extruded_curv.to_pyvista(dim="all").plot(show_edges=True)
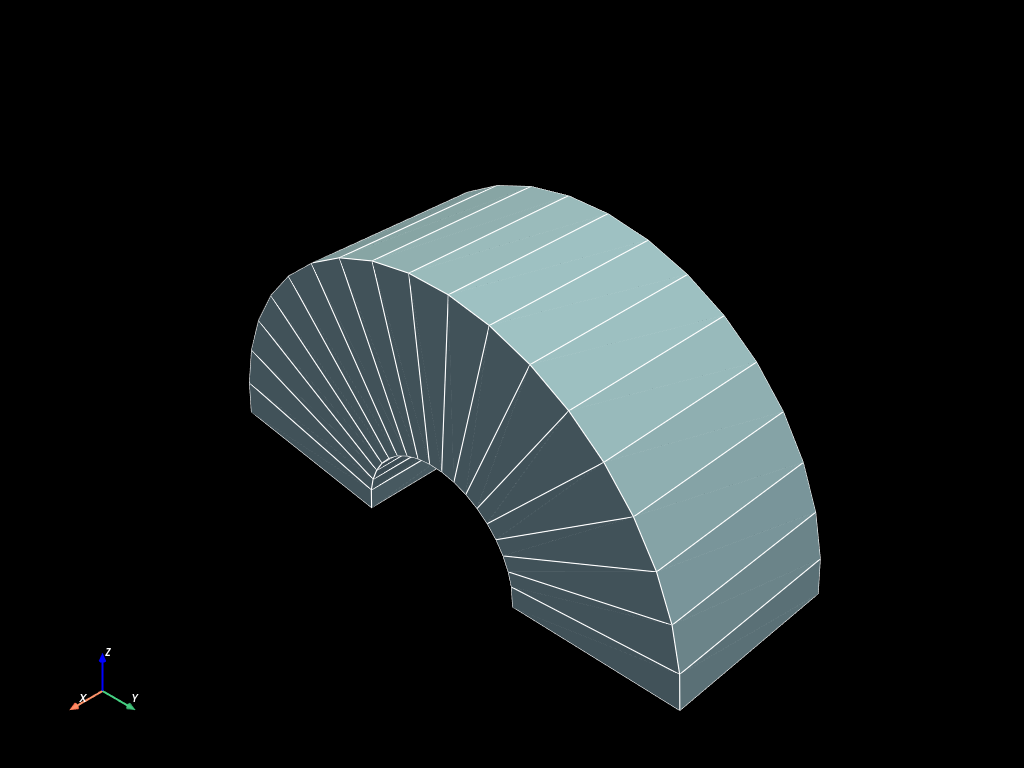

Topological tools
import mefikit as mf
import numpy as np
import pyvista as pv
pv.set_plot_theme("dark")
pv.set_jupyter_backend("static")
Submesh functionality
x = range(3)
y = np.linspace(0.0, 3.0, 5, endpoint=True)
z = np.logspace(-0.5, 1, 4, endpoint=True)
volumes = mf.build_cmesh(x, y, z)
Simple descending_mesh
This functionality is able to compute the descending connectivity of the porvided mesh. It can act on elements of dimension 1, 2 or 3.
faces = volumes.descend()
edges = faces.descend()
vertex = edges.descend()
plotter = pv.Plotter(shape=(1, 3))
plotter.subplot(0, 0)
plotter.add_mesh(faces.to_pyvista().shrink(0.8), show_edges=True)
plotter.subplot(0, 1)
plotter.add_mesh(edges.to_pyvista().shrink(0.8))
plotter.subplot(0, 2)
plotter.add_mesh(vertex.to_pyvista())
plotter.show()
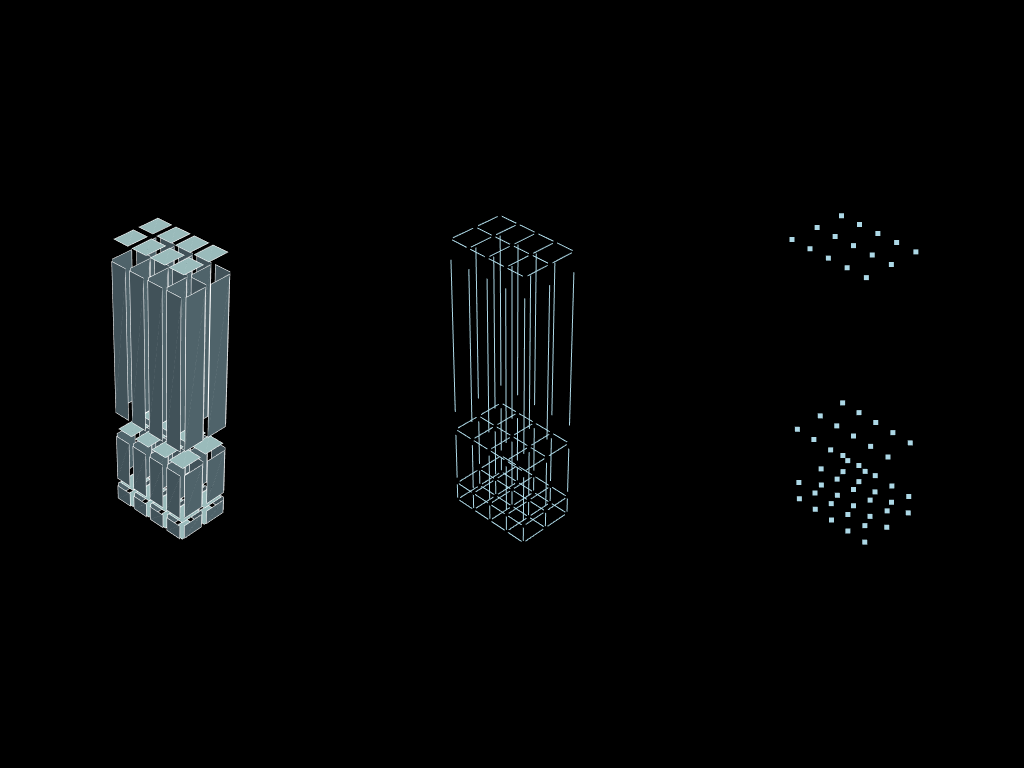

Submesh in one go
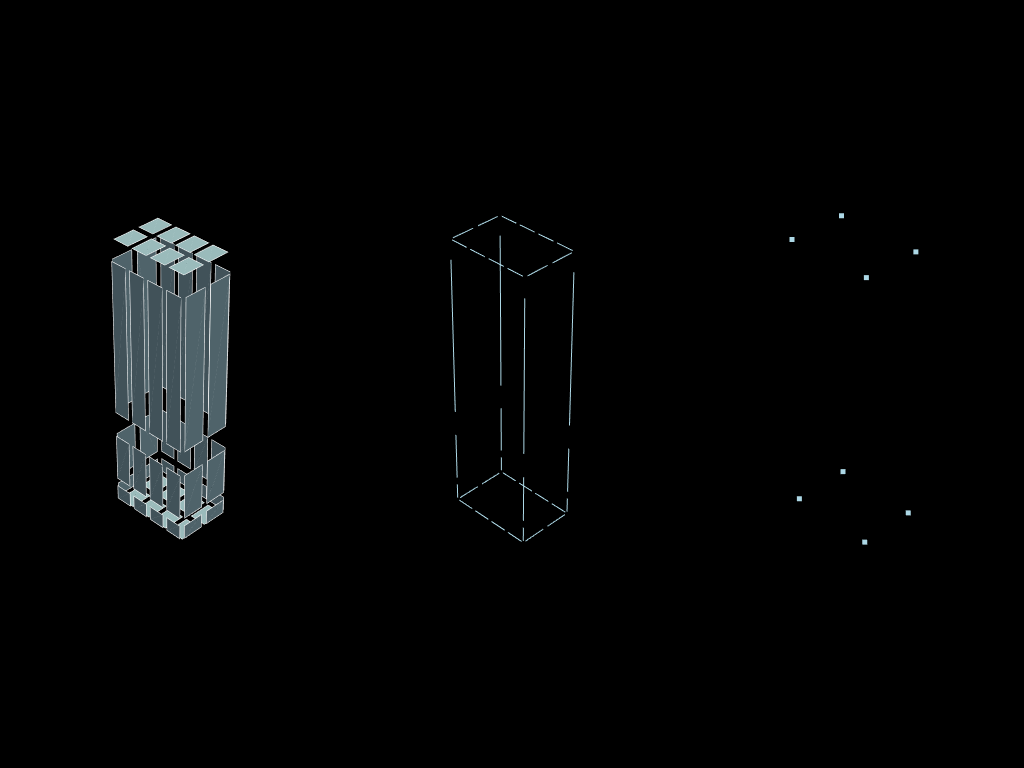
You might want to directly access either the node mesh or the edges mesh. You can ! And going into one step is ever faster than chaining mutliple .descend() calls.
edges = volumes.descend(target_dim=1)
nodes = volumes.descend(target_dim=0)
plotter = pv.Plotter(shape=(1, 2))
plotter.subplot(0, 0)
plotter.add_mesh(edges.to_pyvista().shrink(0.8))
plotter.subplot(0, 1)
plotter.add_mesh(vertex.to_pyvista())
plotter.show()
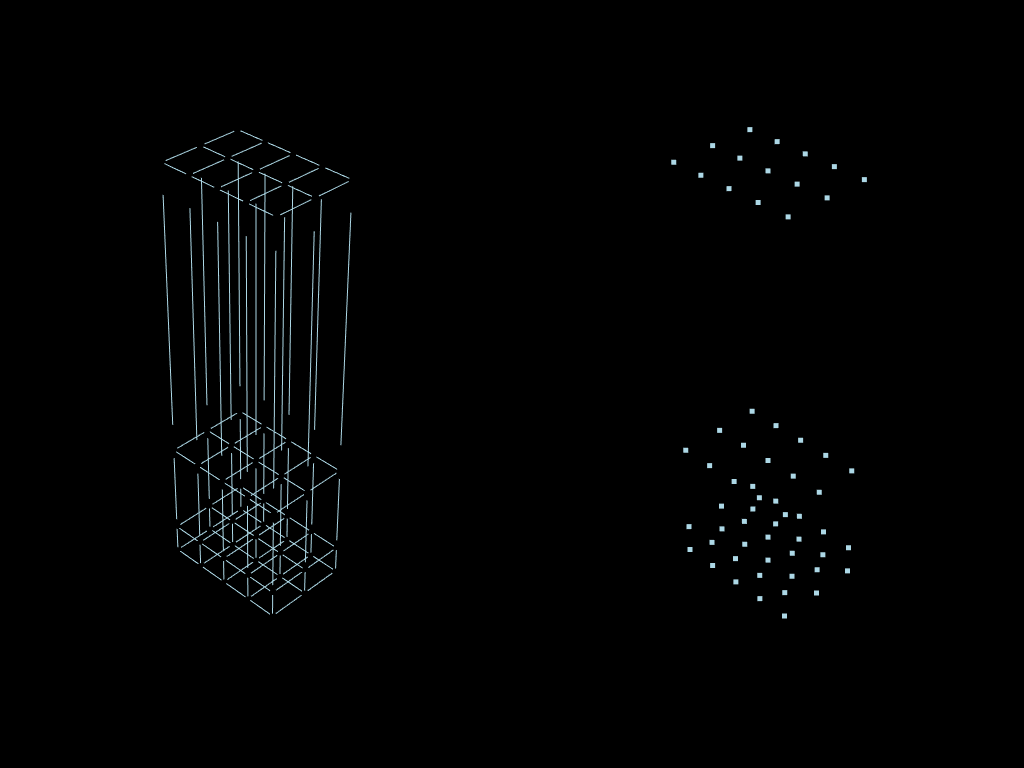
Boundaries computation
As it is very common to compute boundaries on a mesh (for boundary counditions for ex), there is a custom boundaries computation method.
face_bounds = volumes.boundaries()
edge_bounds = volumes.boundaries(target_dim=1)
vertex_bounds = volumes.boundaries(target_dim=0)
plotter = pv.Plotter(shape=(1, 3))
plotter.subplot(0, 0)
plotter.add_mesh(face_bounds.to_pyvista().shrink(0.8), show_edges=True)
plotter.subplot(0, 1)
plotter.add_mesh(edge_bounds.to_pyvista().shrink(0.8))
plotter.subplot(0, 2)
plotter.add_mesh(vertex_bounds.to_pyvista())
plotter.show()

Descend / boundaries update
You can directly update the mesh inplace when computing the descending mesh or the boundaries mesh.
volumes.boundaries_update()
volumes.boundaries_update(target_dim=1)
volumes.to_pyvista().shrink(0.8).plot(show_edges=True)
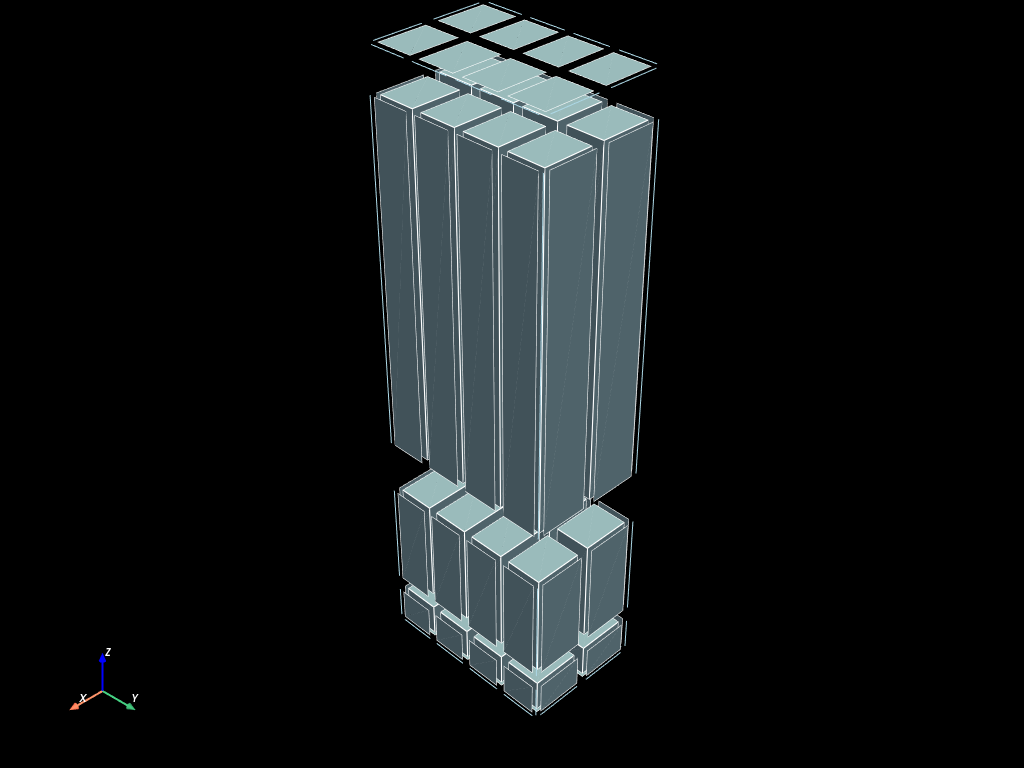

When using the _update version, the elements of the same dimension of the generated mesh are returned as a new mesh.
old_face_mesh = volumes.descend_update()
volumes.to_pyvista().shrink(0.8).plot(show_edges=True)
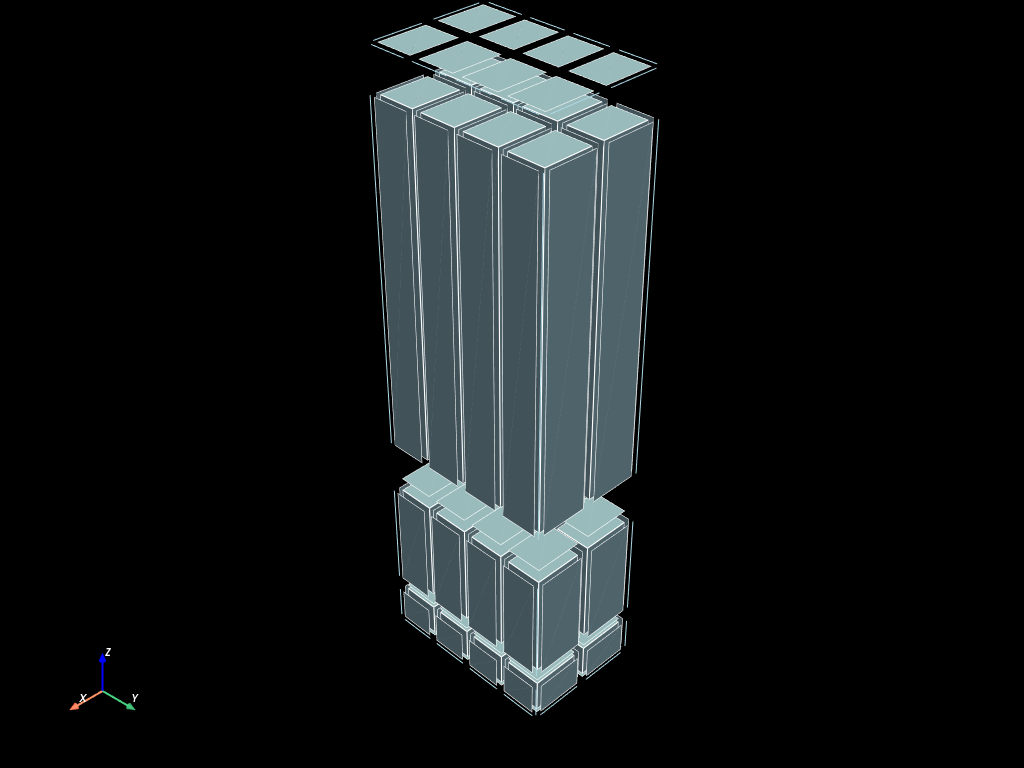

old_face_mesh.to_pyvista().shrink(0.8).plot(show_edges=True)
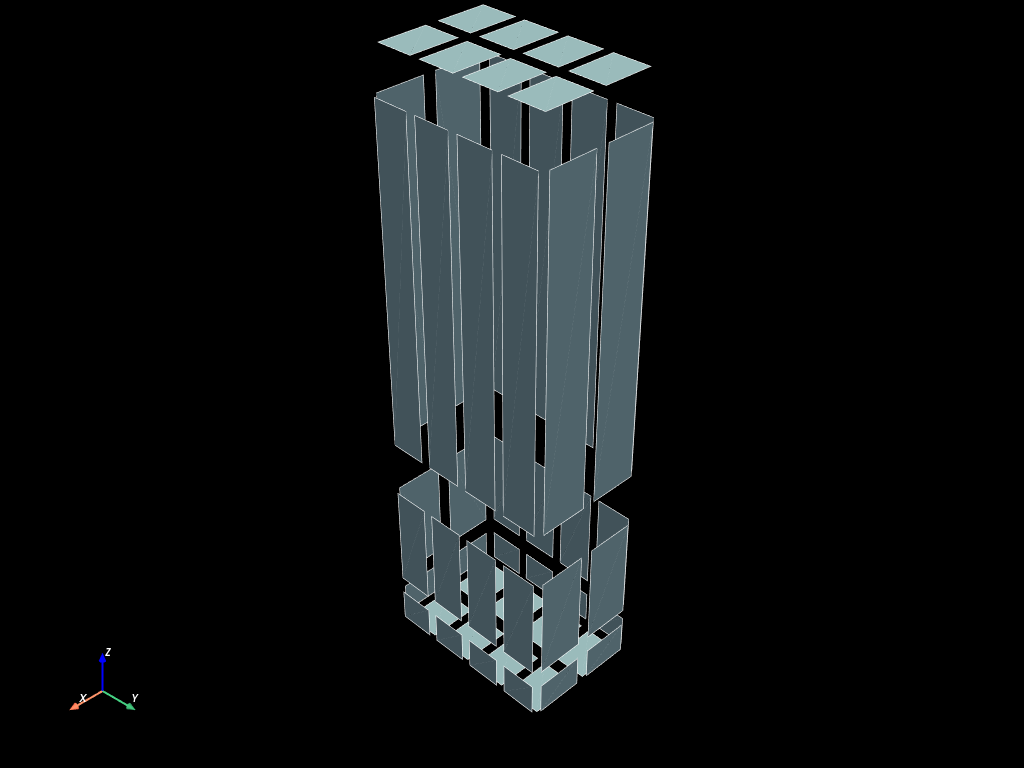

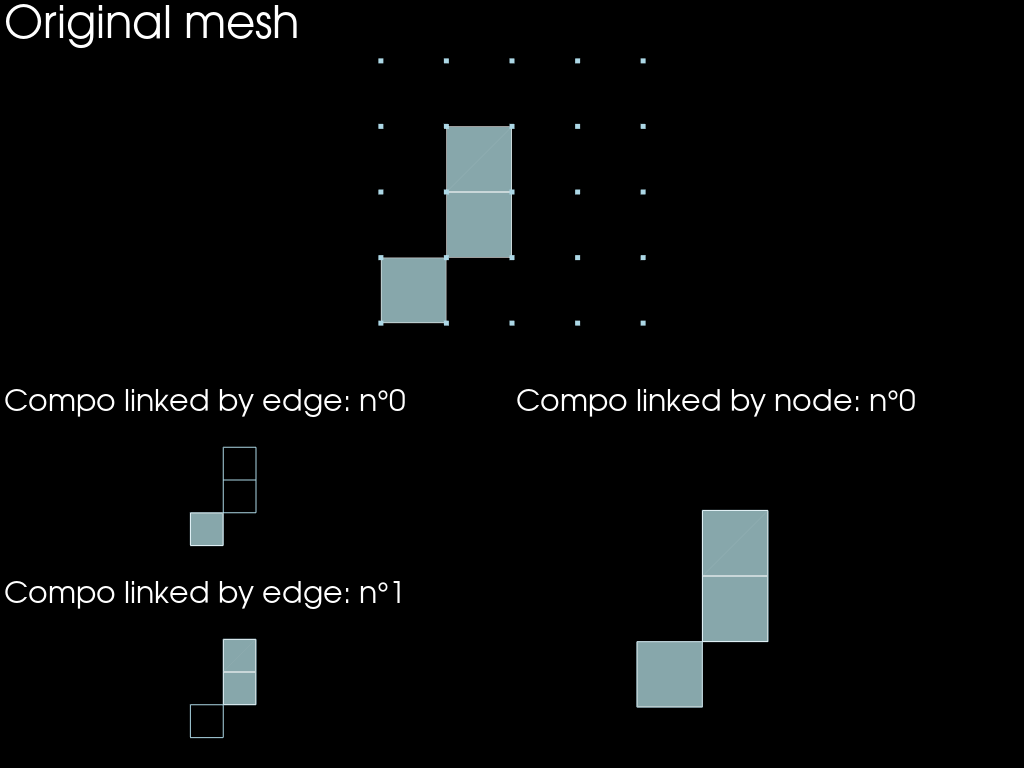
Connected components
x, y = np.meshgrid(np.linspace(0.0, 1.0, 5), np.linspace(0.0, 1.0, 5))
coords = np.c_[x.flatten(), y.flatten()]
conn = np.array(
[
[0, 1, 6, 5],
[6, 7, 12, 11],
# [2, 3, 8, 7],
[11, 12, 17, 16],
],
dtype=np.uint,
)
mesh = mf.UMesh(coords)
mesh.add_regular_block("QUAD4", conn)
mesh.add_regular_block("VERTEX", np.arange(len(coords), dtype=np.uint)[..., np.newaxis])
compos_link_edge = mesh.connected_components(link_dim=1)
compos_link_node = mesh.connected_components(link_dim=0)
print(f"{len(compos_link_edge)=}")
print(f"{len(compos_link_node)=}")
len(compos_link_edge)=2
len(compos_link_node)=1
edges = mesh.descend()
shape = (3, 2)
row_weights = [1.0, 0.5, 0.5]
groups = [
(0, np.s_[:]),
(1, 0),
(2, 0),
(np.s_[1:], 1),
]
plotter = pv.Plotter(shape=shape, groups=groups, row_weights=row_weights)
plotter.subplot(0, 0)
plotter.add_text("Original mesh")
plotter.add_mesh(mesh.to_pyvista(), show_edges=True)
plotter.camera_position = "xy"
for i, compo in enumerate(compos_link_edge):
plotter.subplot(i + 1, 0)
plotter.add_text(f"Compo linked by edge: n°{i}")
plotter.add_mesh(edges.to_pyvista())
plotter.add_mesh(compo.to_pyvista(), show_edges=True)
plotter.camera_position = "xy"
for i, compo in enumerate(compos_link_node):
plotter.subplot(i + 1, 1)
plotter.add_text(f"Compo linked by node: n°{i}")
plotter.add_mesh(edges.to_pyvista())
plotter.add_mesh(compo.to_pyvista(), show_edges=True)
plotter.camera_position = "xy"
plotter.show()
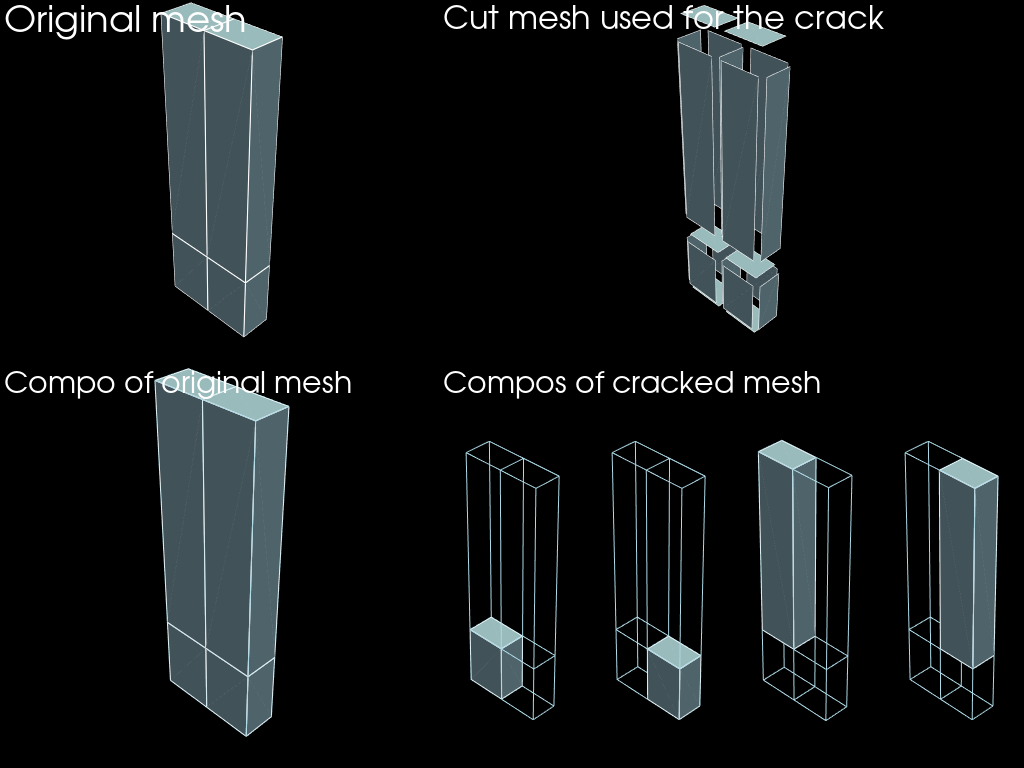
Crack
This feature is the contrary of the merge_nodes feature. It duplicates nodes such that the resulting mesh does not connect on the descending_mesh given.
x = range(2)
y = np.linspace(0.0, 3.0, 3, endpoint=True)
z = np.logspace(0.0, 1.0, 3, endpoint=True)
volumes = mf.build_cmesh(x, y, z)
faces = volumes.descend()
cracked = volumes.crack(faces)
edges = faces.descend()
compos_original = volumes.connected_components()
compos_cracked = cracked.connected_components()
assert len(compos_original) == 1
n_compos = len(compos_cracked)
shape = (3, n_compos + 1)
groups = [
(0, 0),
(0, np.s_[1:]),
(np.s_[1:], 0),
(1, np.s_[1:]),
*((2, i + 1) for i in range(n_compos)),
]
row_weights = [1.0, 0.1, 1.0]
col_weights = [1.5, *(0.5,) * n_compos]
pv.set_jupyter_backend("static")
plotter = pv.Plotter(
shape=shape, groups=groups, row_weights=row_weights, col_weights=col_weights
)
plotter.subplot(0, 0)
plotter.add_text("Original mesh")
plotter.add_mesh(volumes.to_pyvista(), show_edges=True)
plotter.subplot(0, 1)
plotter.add_text("Cut mesh used for the crack")
plotter.add_mesh(faces.to_pyvista().shrink(0.8), show_edges=True)
plotter.subplot(1, 0)
plotter.add_text("Compo of original mesh")
plotter.add_mesh(edges.to_pyvista())
plotter.add_mesh(compos_original[0].to_pyvista(), show_edges=True)
plotter.subplot(1, 1)
plotter.add_text("Compos of cracked mesh")
for i, compo in enumerate(compos_cracked):
plotter.subplot(2, i + 1)
plotter.add_mesh(edges.to_pyvista())
plotter.add_mesh(compo.to_pyvista(), show_edges=True)
plotter.camera.zoom(2)
plotter.show()
Geometrical tools
import mefikit as mf
import numpy as np
import pyvista as pv
pv.set_plot_theme("dark")
pv.set_jupyter_backend("static")
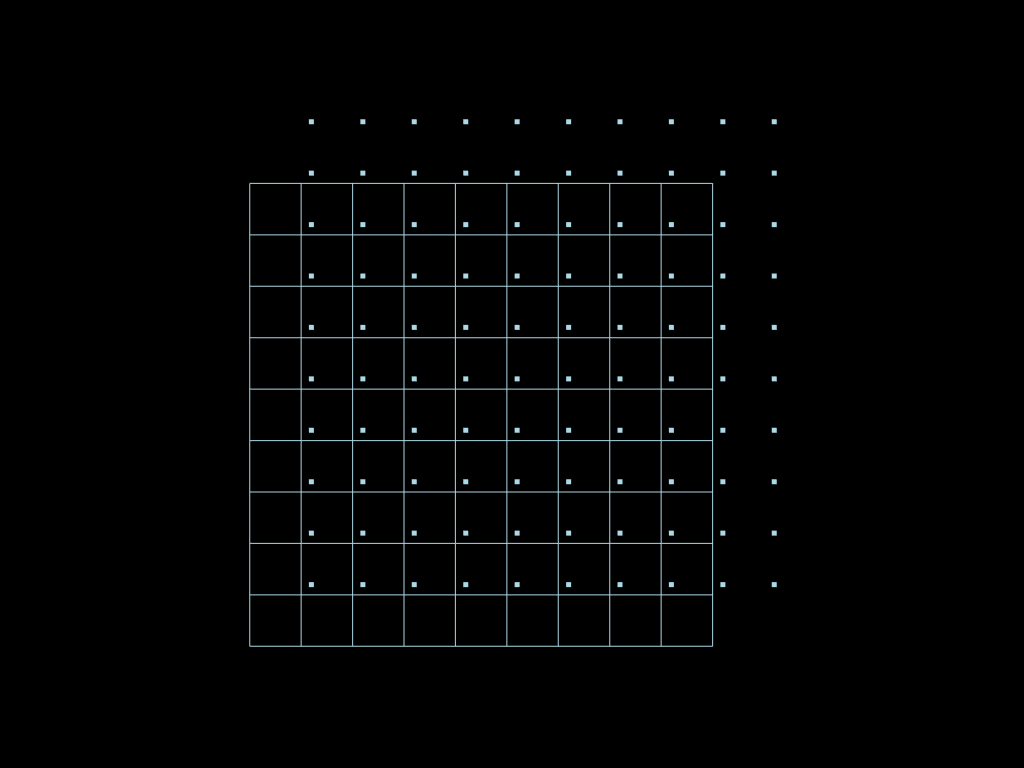
Snap points
x = np.linspace(0.0, 3.0, 10, endpoint=True)
mesh = mf.build_cmesh(x, x)
eps = 0.1
dec = x[-1] / len(x) + eps
x2 = np.linspace(dec, x[-1] + dec, len(x), endpoint=True)
mesh2 = mf.build_cmesh(x2, x2)
Note: epsilon value used in the following operation is big enough so that there are multiple candidates for some points, but low enough so that there is no degenerated cell created.
snaped = mesh.snap(mesh2, eps=x[-1] / len(x))
pt = pv.Plotter()
pt.add_mesh(mesh.descend().to_pyvista())
pt.add_mesh(mesh2.descend(target_dim=0).to_pyvista())
pt.show(cpos="xy")
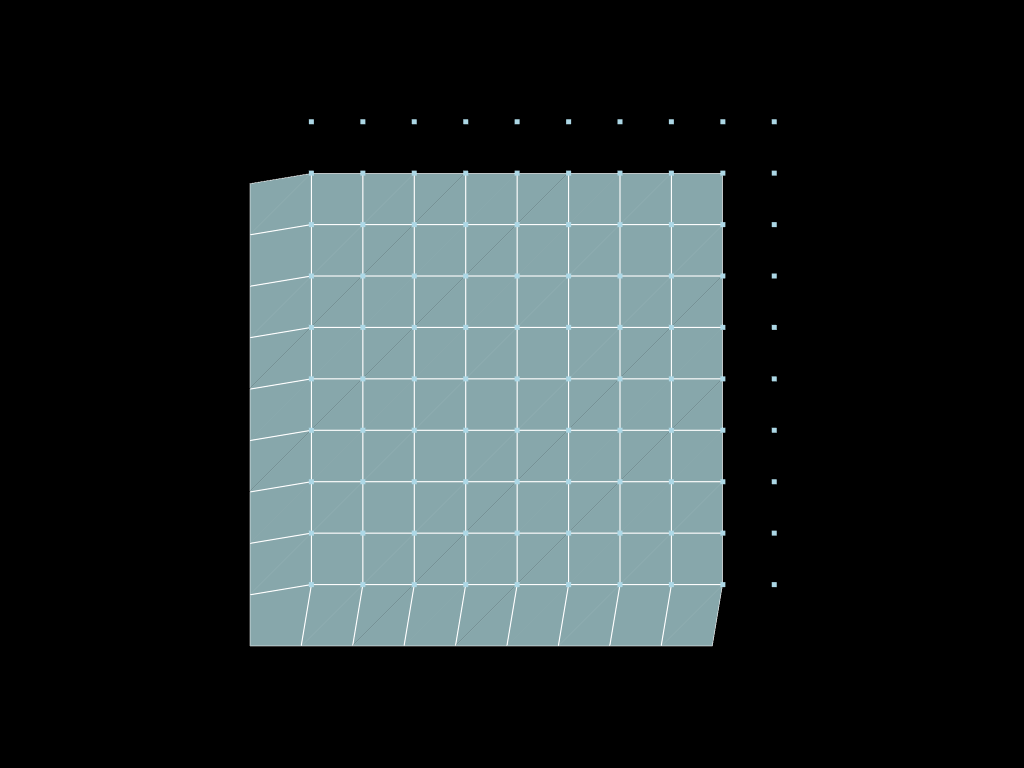
pt = pv.Plotter()
pt.add_mesh(snaped.to_pyvista(), show_edges=True)
pt.add_mesh(mesh2.descend(target_dim=0).to_pyvista())
pt.show(cpos="xy")
Merge nodes
x = range(2)
y = np.linspace(0.0, 3.0, 3, endpoint=True)
z = np.logspace(0.0, 1.0, 3, endpoint=True)
volumes = mf.build_cmesh(x, y, z)
faces = volumes.descend()
cracked = volumes.crack(faces)
merged = cracked.merge_nodes()
edges = faces.descend()
compos_merged = merged.connected_components()
compos_cracked = cracked.connected_components()
assert len(compos_merged) == 1
n_compos = len(compos_cracked)
shape = (3, n_compos + 1)
groups = [
(0, np.s_[:-1]), # cracked
(0, n_compos), # merged
(1, np.s_[:-1]), # cracked txt
(np.s_[1:], n_compos), # merged compos
*((2, i) for i in range(n_compos)), # cracked compos
]
row_weights = [1.0, 0.1, 1.0]
col_weights = [*(0.5,) * n_compos, 1.5]
pv.set_jupyter_backend("static")
plotter = pv.Plotter(
shape=shape, groups=groups, row_weights=row_weights, col_weights=col_weights
)
plotter.subplot(0, n_compos)
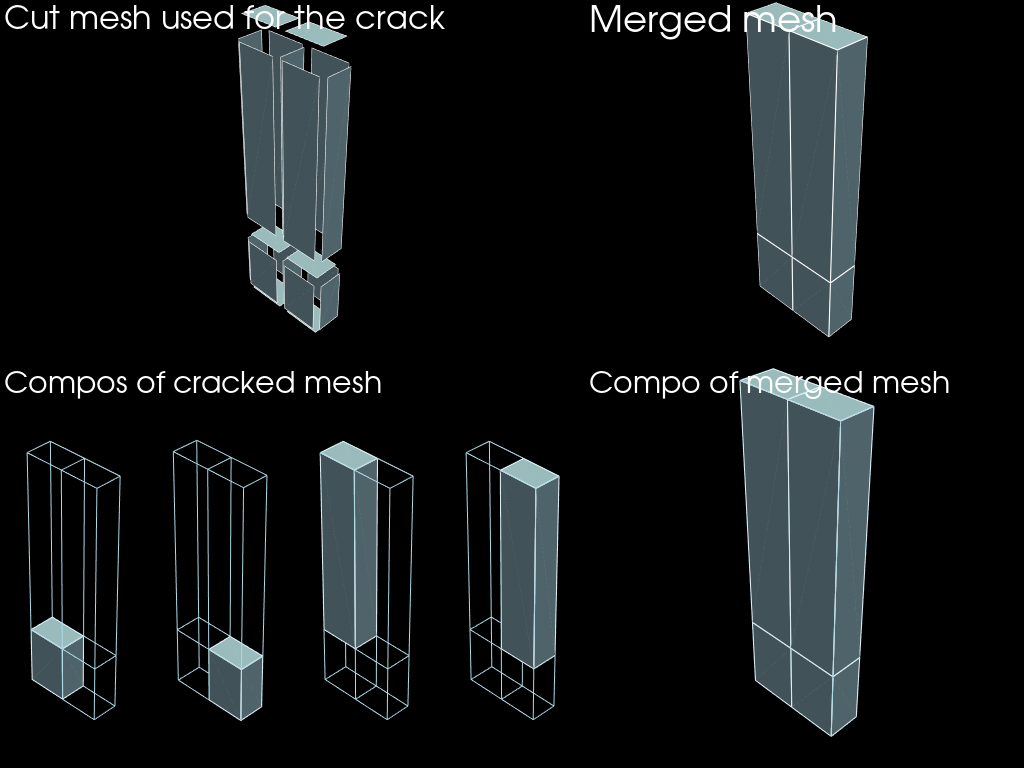
plotter.add_text("Merged mesh")
plotter.add_mesh(merged.to_pyvista(), show_edges=True)
plotter.subplot(0, 0)
plotter.add_text("Cut mesh used for the crack")
plotter.add_mesh(faces.to_pyvista().shrink(0.8), show_edges=True)
plotter.subplot(1, n_compos)
plotter.add_text("Compo of merged mesh")
plotter.add_mesh(edges.to_pyvista())
plotter.add_mesh(compos_merged[0].to_pyvista(), show_edges=True)
plotter.subplot(1, 0)
plotter.add_text("Compos of cracked mesh")
for i, compo in enumerate(compos_cracked):
plotter.subplot(2, i)
plotter.add_mesh(edges.to_pyvista())
plotter.add_mesh(compo.to_pyvista(), show_edges=True)
plotter.camera.zoom(2)
plotter.show()
Selection tool
import mefikit as mf
import numpy as np
import pyvista as pv
pv.set_plot_theme("dark")
pv.set_jupyter_backend("static")
Element selection
Elements can be selected based on there:
- types
- ids
- dimensions
Elements can be selected based on there centroid position. The selection methods using centroids are :
- bbox
- sphere
- rectangle
- circle
As you can guess, bbox and sphere should be used for 3d meshes and rectangle and circle for 2d meshes.
Elements can be selected based on the nodes position and a boolean : whether to select element with all nodes matching the coundition or any node matching the coundition.
- nbbox
- nsphere
- nrect
- ncircle
- nids
Elements can be selected based on their group appartenance :
- inside
- outside
Elements can be selected based on their scalar fields values :
- FieldExpr > FieldExpr
- FieldExpr >= FieldExpr
- FieldExpr < FieldExpr
- FieldExpr <= FieldExpr
- FieldExpr == FieldExpr
The Selection type
Here the types manipulated are Selection. They are simple objects that know how to compose themselves.
clip = mf.sel.bbox([-np.inf, -np.inf, -np.inf], [np.inf, np.inf, 0.5])
sphere = mf.sel.sphere([0.5, 0.5, 0.5], 0.5)
x = np.linspace(0.0, 1.0, 20, endpoint=True)
volumes = mf.build_cmesh(x, x, x)
volumes.select(clip).to_pyvista().plot()
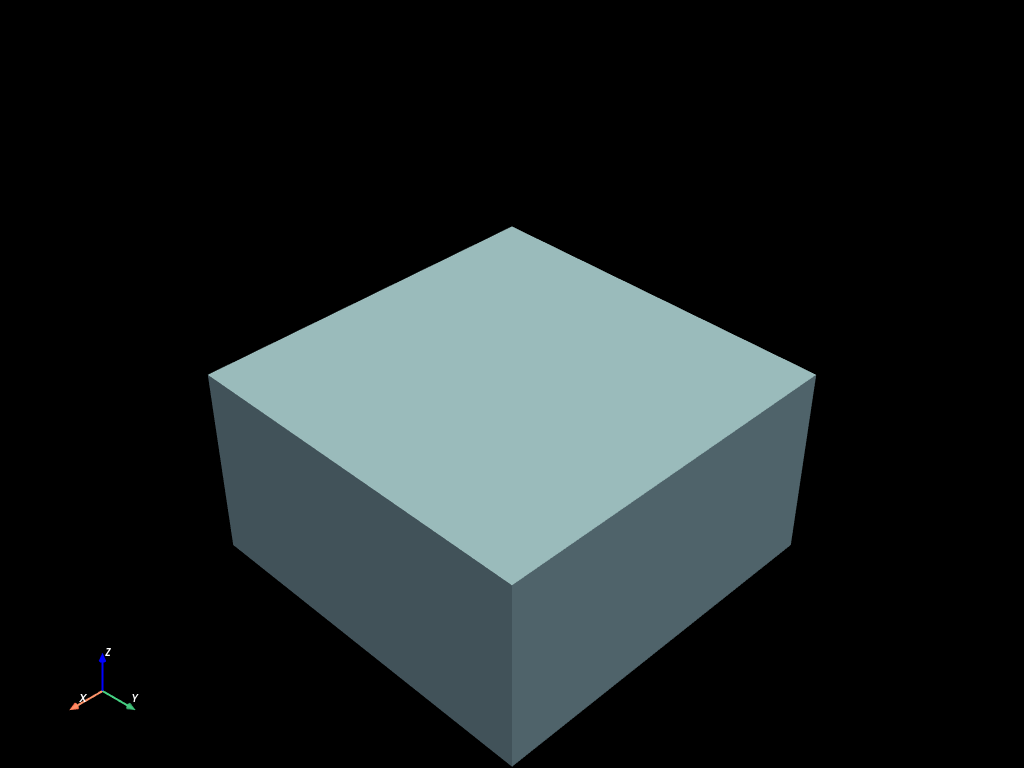

volumes.select(sphere).to_pyvista().plot()

Selection composition
One of the great strength of the select method is its composability ! Whatch by yourself.
The operators &, |, ^, - and ~ are available.
x = np.linspace(0.0, 2.0, 100)
y = np.linspace(0.0, 1.0, 50)
faces = mf.build_cmesh(x, y)
circle1 = mf.sel.circle([0.75, 0.5], 0.5)
circle2 = mf.sel.circle([1.25, 0.5], 0.5)
union = faces.select(circle1 | circle2)
pt = pv.Plotter()
pt.add_mesh(faces.descend().to_pyvista())
pt.add_mesh(union.to_pyvista())
pt.camera_position = "xy"
pt.show()
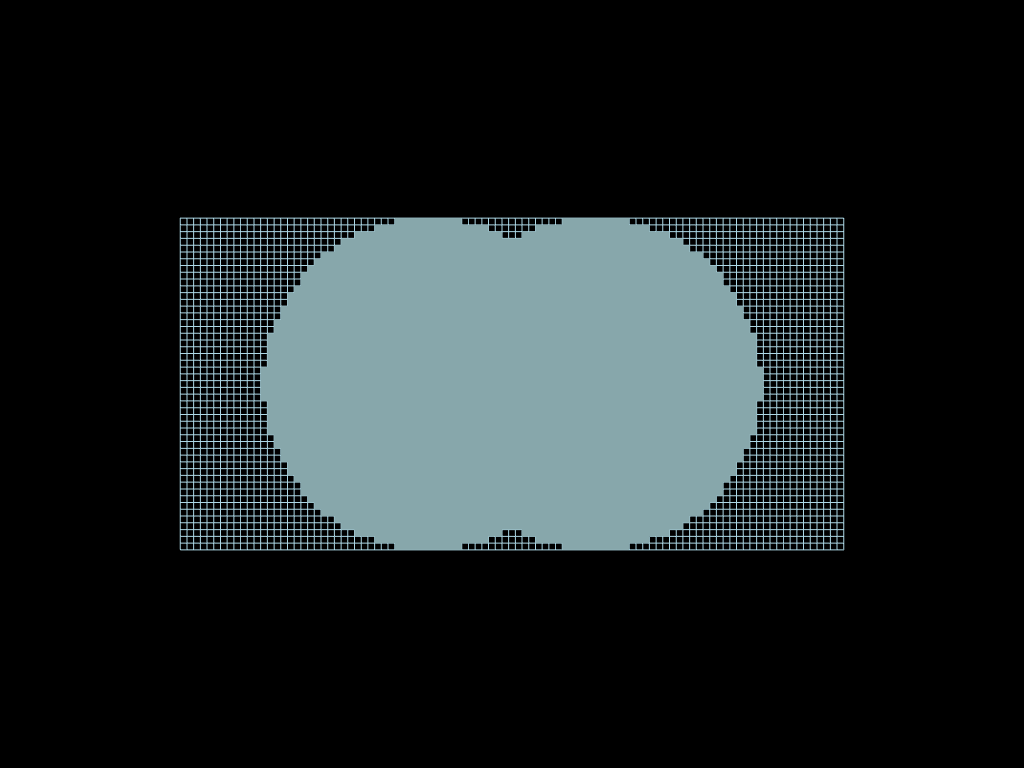

intersection = faces.select(circle1 & circle2)
pt = pv.Plotter()
pt.add_mesh(faces.descend().to_pyvista())
pt.add_mesh(intersection.to_pyvista())
pt.camera_position = "xy"
pt.show()
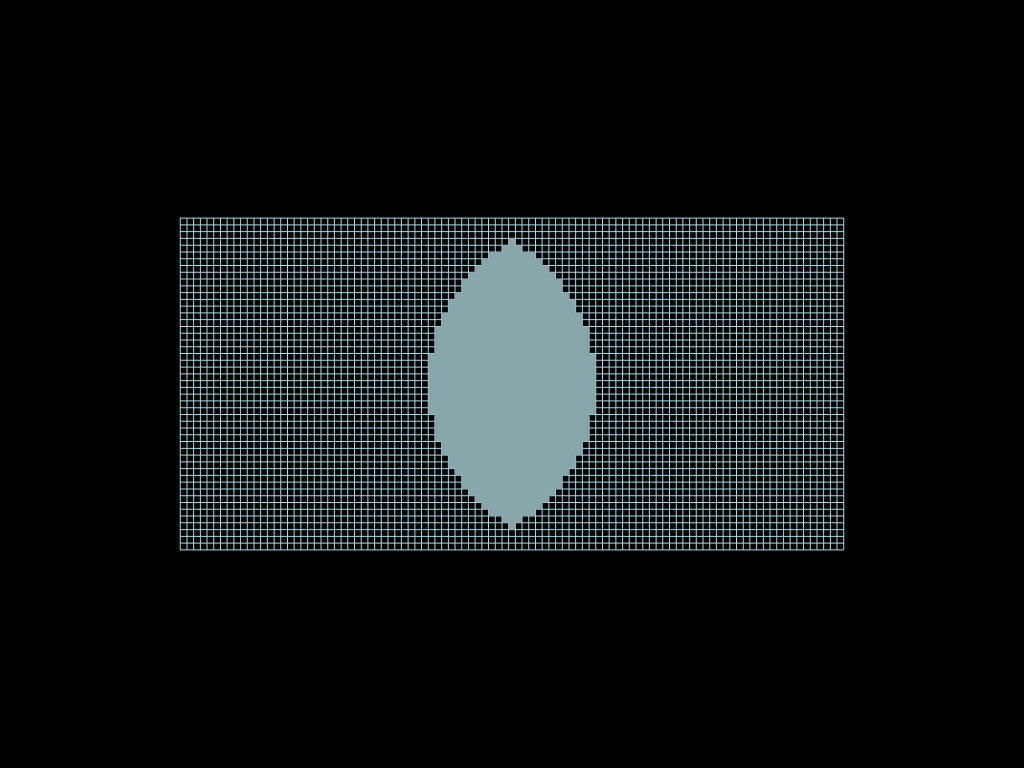

sym_diff = faces.select(circle1 ^ circle2)
pt = pv.Plotter()
pt.add_mesh(faces.descend().to_pyvista())
pt.add_mesh(sym_diff.to_pyvista())
pt.camera_position = "xy"
pt.show()
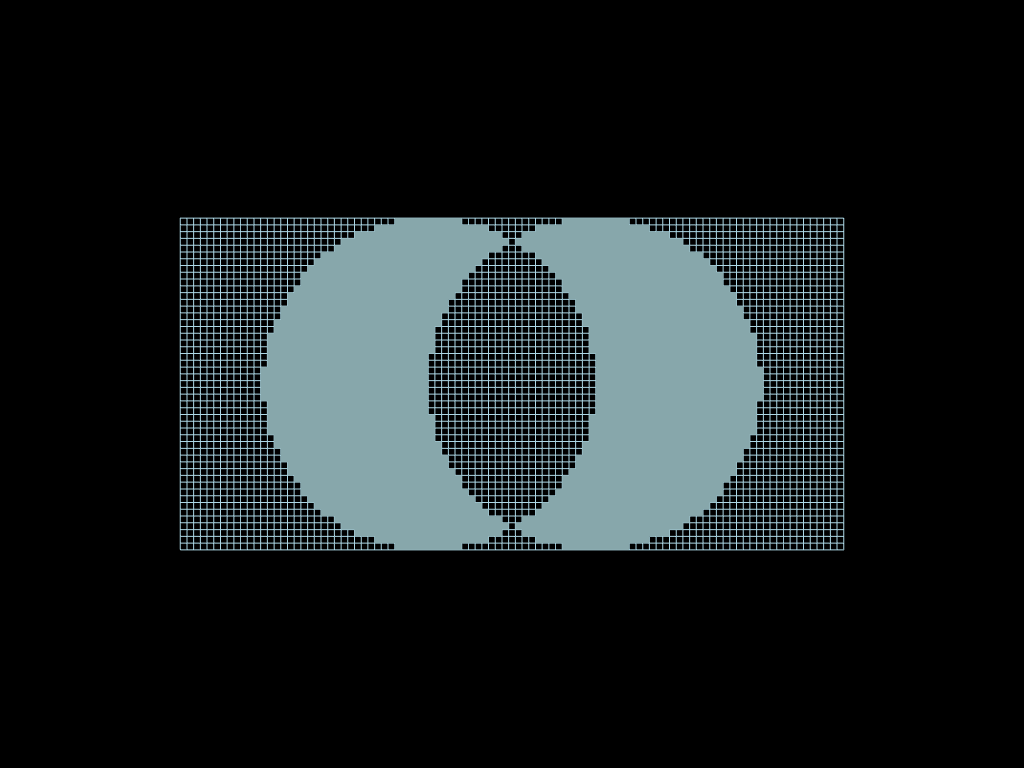

diff = faces.select(circle1 - circle2)
pt = pv.Plotter()
pt.add_mesh(faces.descend().to_pyvista())
pt.add_mesh(diff.to_pyvista())
pt.camera_position = "xy"
pt.show()
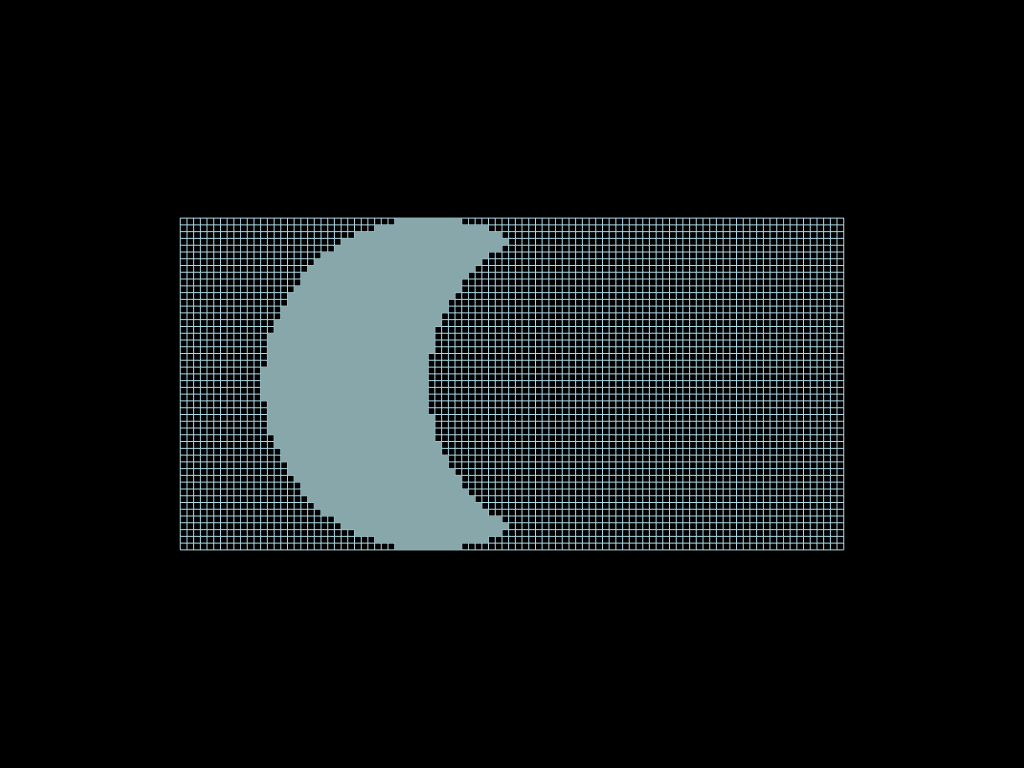

notsel = faces.select(~circle1)
pt = pv.Plotter()
pt.add_mesh(faces.descend().to_pyvista())
pt.add_mesh(notsel.to_pyvista())
pt.camera_position = "xy"
pt.show()
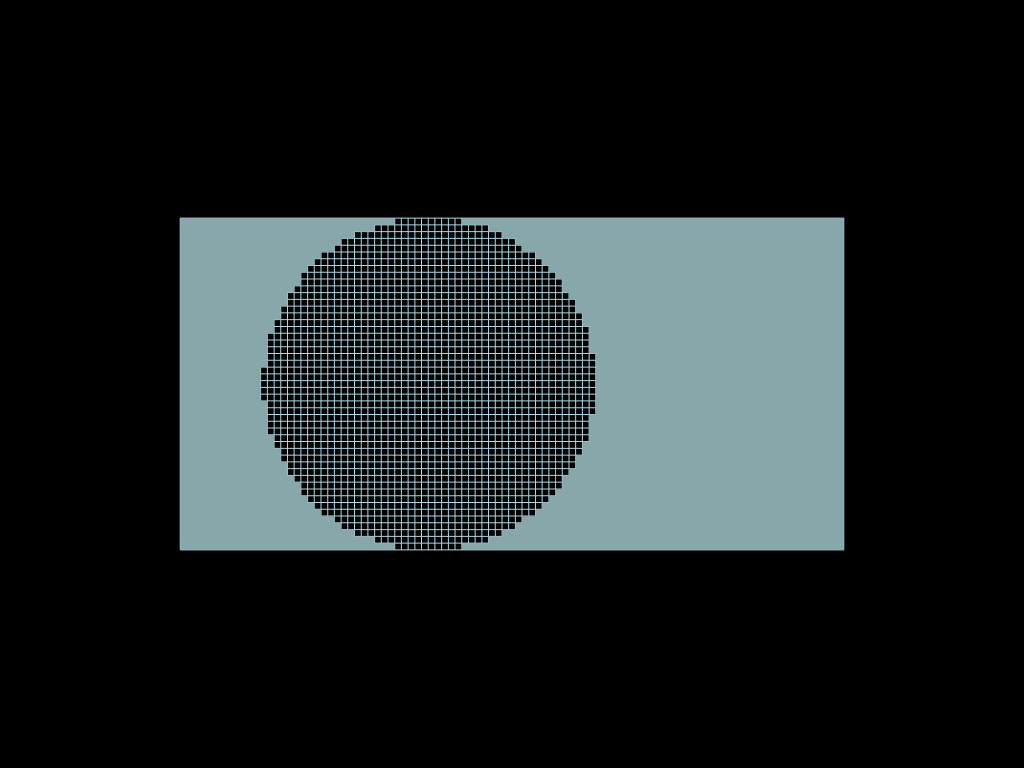

A 3D complex example
sphere = mf.sel.sphere([0.5, 0.5, 0.5], 0.5)
clip_x = mf.sel.bbox([0.5, -np.inf, -np.inf], [np.inf, np.inf, np.inf]) # x > 0.5
clip_z = mf.sel.bbox([-np.inf, -np.inf, -np.inf], [np.inf, np.inf, 0.5]) # z < 0.5
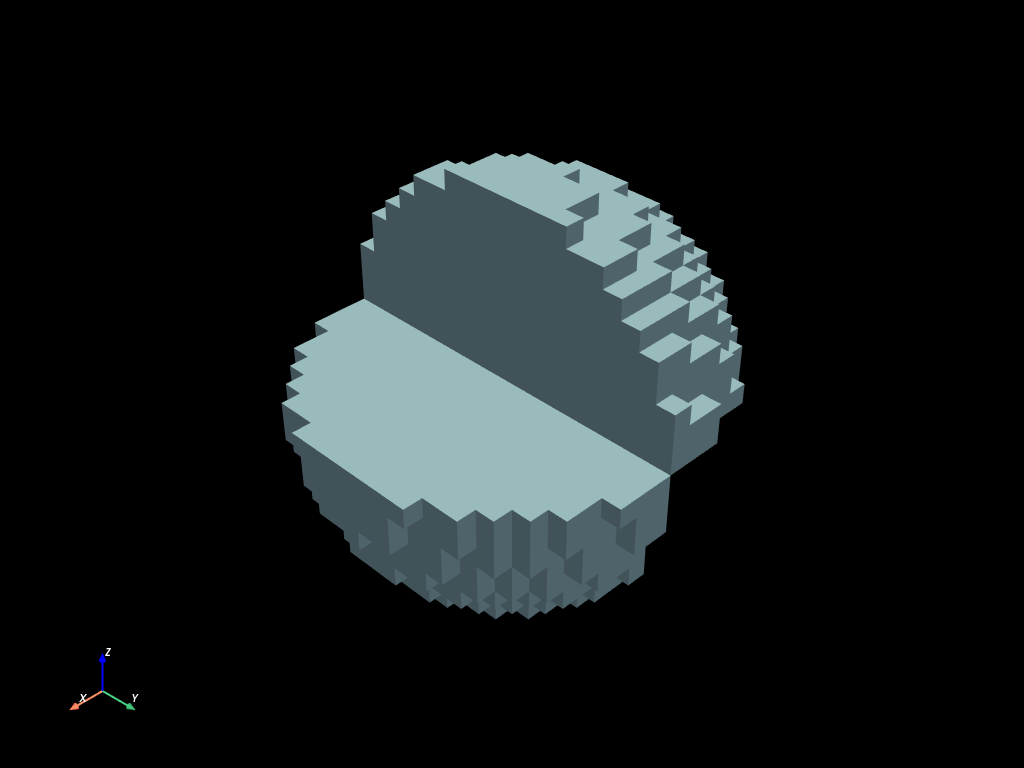
volumes.select(
(clip_x & sphere & clip_z) | (sphere & ~clip_x & ~clip_z)
).to_pyvista().plot()

How does it works ?
Each objects generated by a selection function (a function from the mf.sel module) is of the Selection type. It implements operators so that it knows how to compose in an expression. That way an expression generates a new Selection which can be interpreted by the .select(expr) method.
print(sphere)
CentroidSelection(
Sphere {
center: [
0.5,
0.5,
0.5,
],
r2: 0.5,
},
)
print(~(sphere & clip_x))
NotExpr(
NotExpr(
BinarayExpr(
BinarayExpr {
operator: And,
left: CentroidSelection(
Sphere {
center: [
0.5,
0.5,
0.5,
],
r2: 0.5,
},
),
right: CentroidSelection(
BBox {
min: [
0.5,
-inf,
-inf,
],
max: [
inf,
inf,
inf,
],
},
),
},
),
),
)
You can see two layers of NotExpr and BinaryExpr. That is perfectly normal, it does not mean that the operation is applied twice, both NotExpr operations and both BinaryExpr op actually comes from different namespaces and it is just a form of encapsulation (first is a variant, second is an enum).
Fields
import mefikit as mf
import numpy as np
import pyvista as pv
pv.set_plot_theme("dark")
pv.set_jupyter_backend("static")
Field expressions
FieldExpr are composition of floats and mf.sel.field(“fieldname”) or custom fields :
- field(“toto”) * field(“tata”)
- field(“toto”) + field(“tata”)
- field(“toto”) - field(“tata”)
- field(“toto”) / field(“tata”)
- field(“toto”) ** field(“tata”)
- field(“toto”).dot(field(“tata”))
- field(“toto”) @ field(“tata”)
- sin(field(“toto”))
- cos(field(“toto”))
- abs(field(“toto”))
- log(field(“toto”))
- log10(field(“toto”))
- exp(field(“toto”))
- field(“toto”)[0]
- normals()
- x()
- y()
- z()
- centroids()
x = np.logspace(-5, 0.0, 1000)
mesh2 = mf.build_cmesh(x, x)
mesh2.measure_update()
mesh2.to_pyvista().plot()
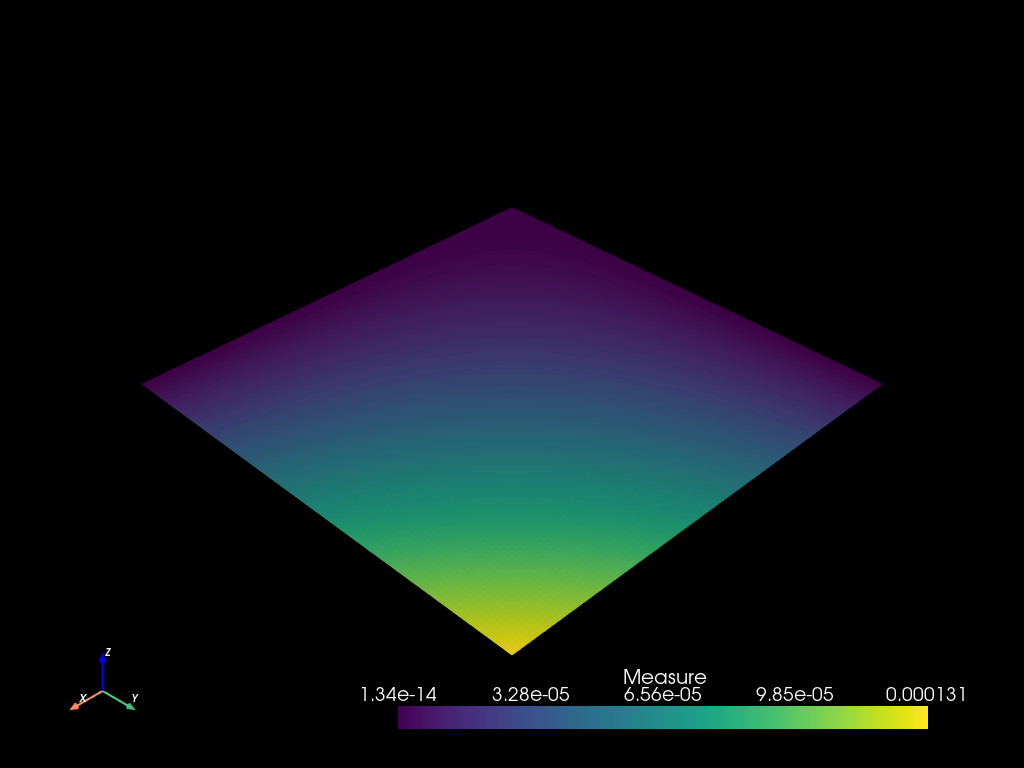
m = mf.Field("Measure")
Field operations evaluation
mesh2.eval_update("4 * M2", m * m * 4.0)
mesh2.to_pyvista().plot()
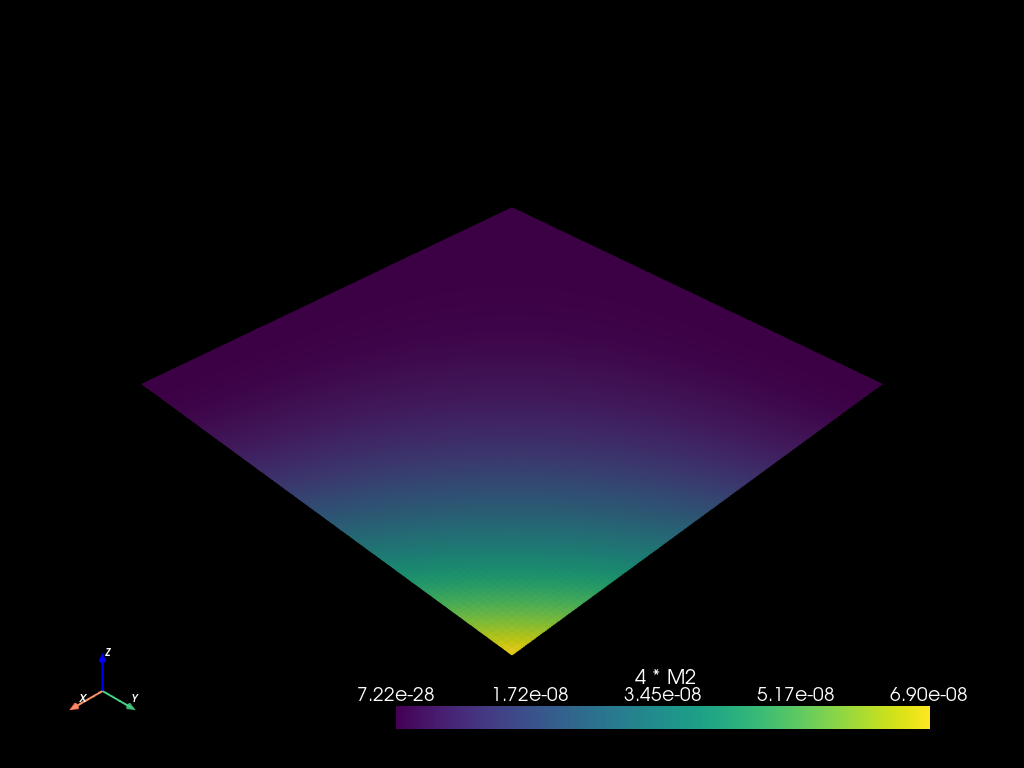
mesh2.fields()
{'4 * M2': {'QUAD4': array([7.22038631e-28, 7.38874101e-28, 7.56102117e-28, ...,
6.58446349e-08, 6.73799065e-08, 6.89509753e-08], shape=(998001,))},
'Measure': {'QUAD4': array([1.34353883e-14, 1.35911194e-14, 1.37486555e-14, ...,
1.28301047e-04, 1.29788199e-04, 1.31292589e-04], shape=(998001,))}}
m2 = mf.Field("4 * M2")
mesh2.eval(m2 - 4.0 * m.square())
{'QUAD4': array([0., 0., 0., ..., 0., 0., 0.], shape=(998001,))}
How does it work ?
print(4.0 * m * m)
BinarayExpr {
operator: Mul,
left: BinarayExpr {
operator: Mul,
left: Array(
4.0, shape=[], strides=[], layout=CFcf (0xf), dynamic ndim=0,
),
right: Field(
"Measure",
),
},
right: Field(
"Measure",
),
}
Why is it awesome ?
This enables two patterns :
- reusability and composition of filters
- evaluation optimizations, some selection filters are evaluated in parallel, some are evaluated first if they are discriminant
Field to Selection
Fields can be converted to threshold selections :
th = (m > 3.25e-5) & (m < 1e-4)
m2sel = mesh2.select(th)
pvm2: pv.UnstructuredGrid = m2sel.to_pyvista()
pvm2.active_scalars_name = "Measure"
pvm2.plot()
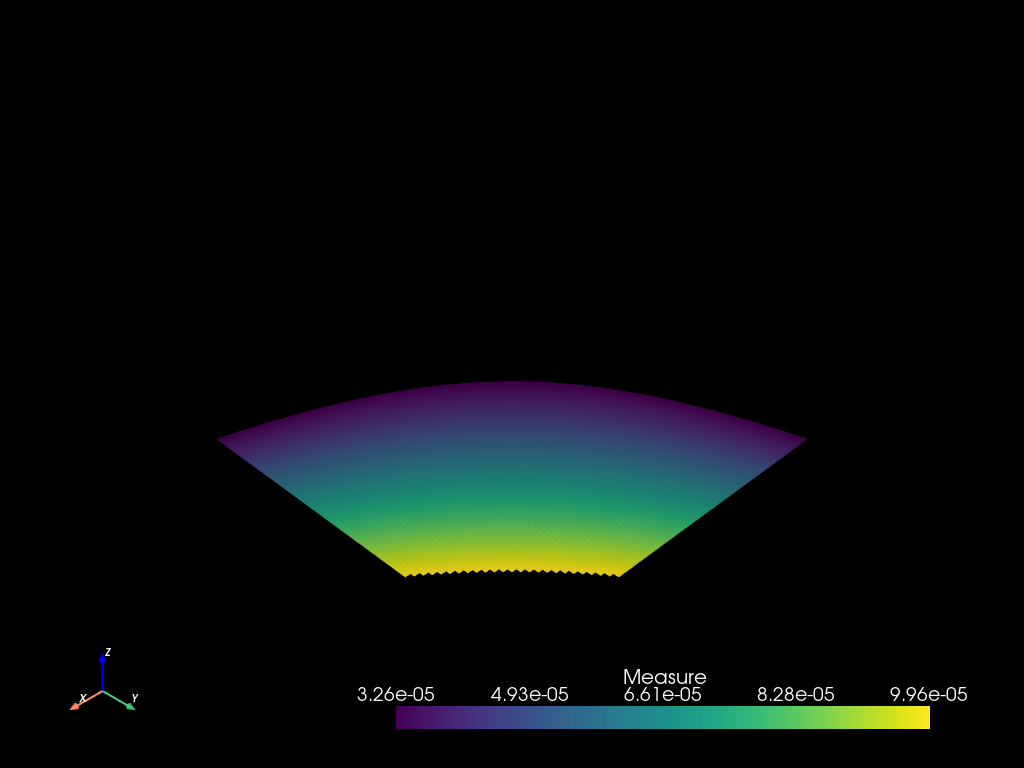
Thoses threasholds selections can be combined with other selections.
r = mf.sel.rect([0.25, 0.25], [0.7, 0.7])
c = mf.sel.circle([0.875, 0.875], 0.05)
mesh2.select((m2 > 4e-9) - r - c).to_pyvista().plot()
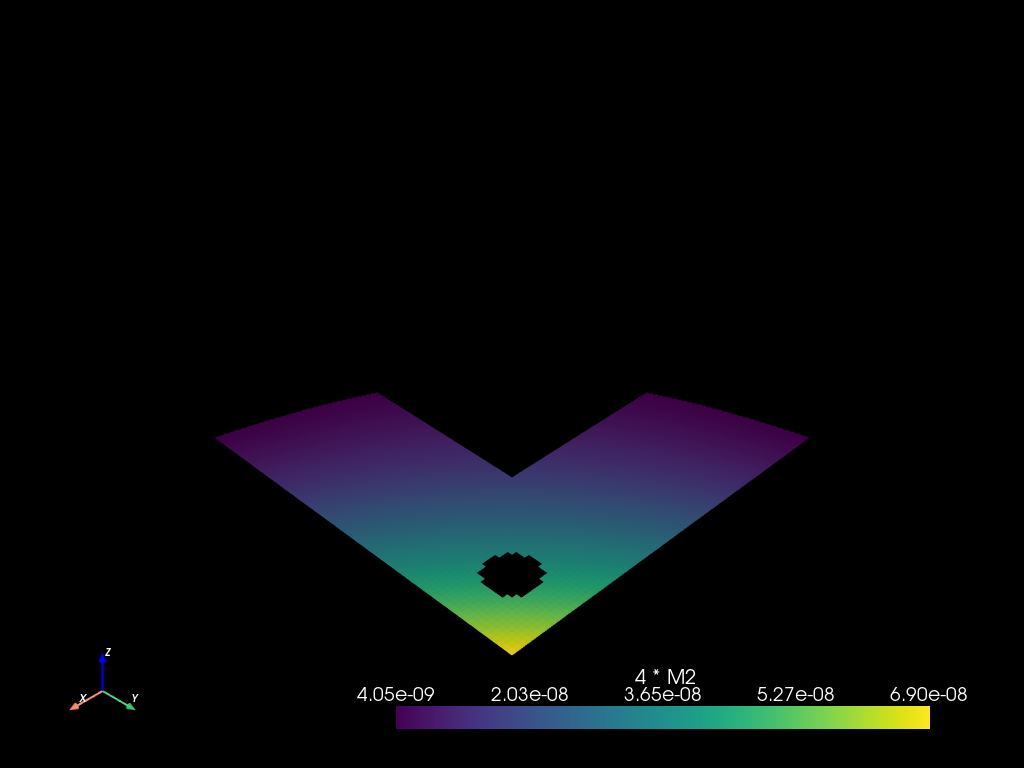
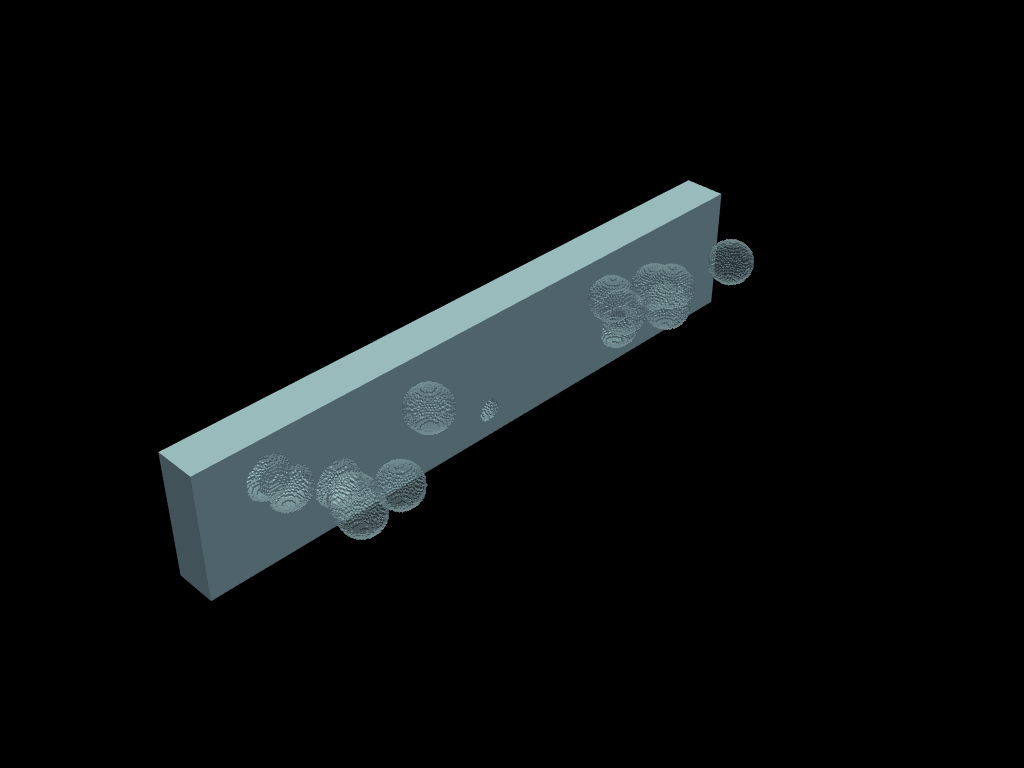
Bubbles example
import mefikit as mf
import numpy as np
import pyvista as pv
rng = np.random.default_rng(seed=123)
pv.set_plot_theme("dark")
pv.set_jupyter_backend("static")
xmax = 5.0
ymax = 1.0
r = 0.17
nb = 15
nr = 12.5
nx = int(xmax / r * nr)
ny = int(ymax / r * nr)
print(f"Number of elements : {nx * ny * ny:,}")
Number of elements : 1,955,743
xc = rng.uniform(r, xmax - r, nb)
yc = rng.uniform(r, ymax - r, nb)
zc = rng.uniform(r, ymax - r, nb)
spheres = [mf.sel.sphere([x, y, z], r) for x, y, z in zip(xc, yc, zc)]
sphere_union = spheres[0]
for s in spheres[1:]:
sphere_union = sphere_union | s
x = np.linspace(0.0, xmax, nx)
y = np.linspace(0.0, ymax, ny)
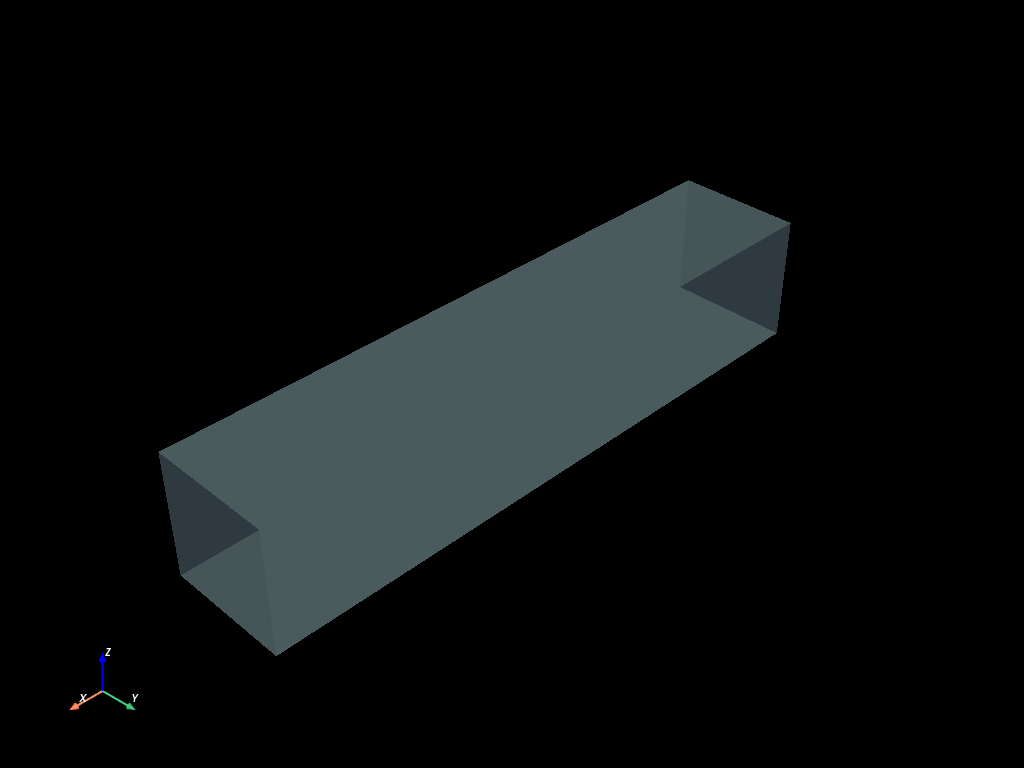
volumes = mf.build_cmesh(x, y, y)
volumes.boundaries().to_pyvista().plot(opacity=0.4)

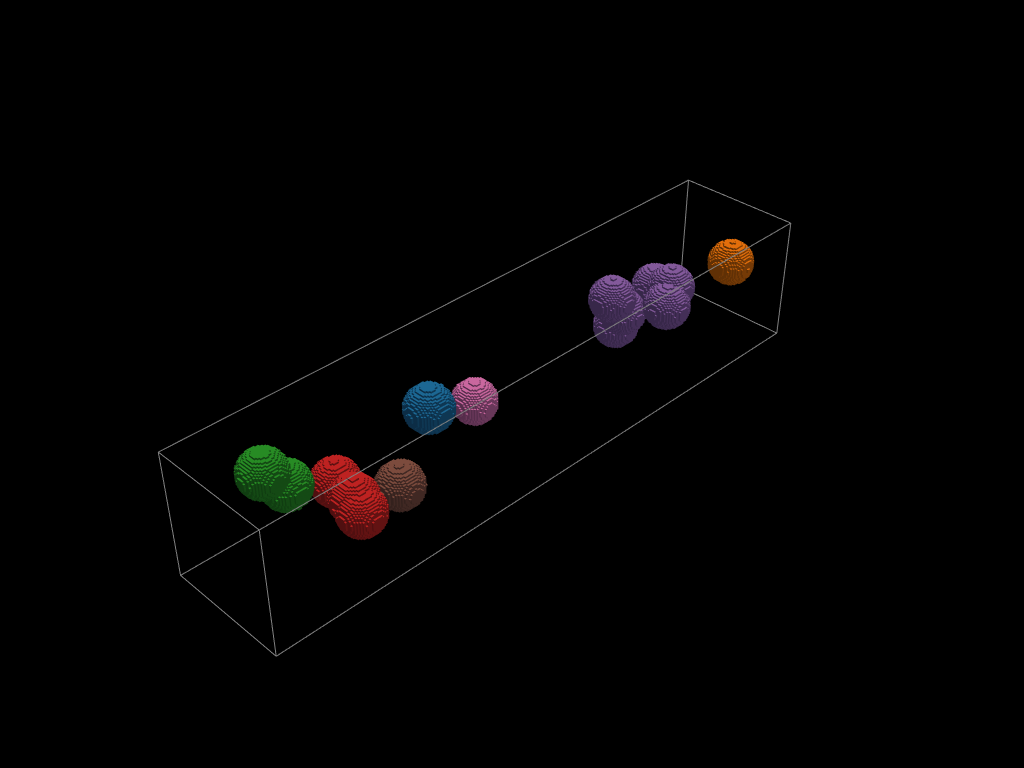
inner_bubbles = volumes.select(sphere_union)
interface = inner_bubbles.boundaries()
cracked = volumes.crack(interface)
cracked.boundaries().to_pyvista().plot(opacity=0.4)
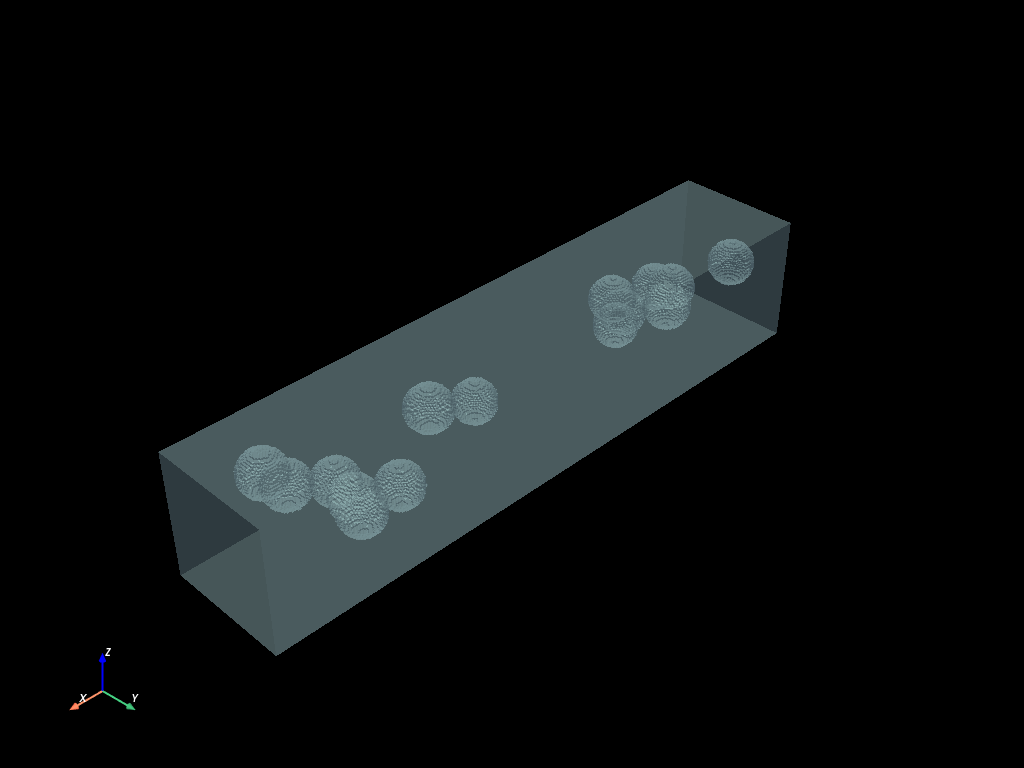
bubble_groups = inner_bubbles.connected_components()
pv.global_theme.color_cycler = "default"
pl = pv.Plotter()
for c in bubble_groups:
compo = c.to_pyvista()
pl.add_mesh(compo)
pl.add_mesh(volumes.boundaries(target_dim=1).to_pyvista())
pl.show()
pv.global_theme.color_cycler = None

clip1 = mf.sel.bbox([-np.inf] * 3, [np.inf, ymax / 3.0, np.inf])
pl = pv.Plotter()
pl.add_mesh(volumes.select(clip1 & ~sphere_union).to_pyvista())
pl.add_mesh(interface.to_pyvista(), opacity=0.4)
pl.show()
